The liver is the largest internal organ and it provides many essential metabolic, exocrine and endocrine functions. Hepatocytes are the principal cell type in the liver and these along with biliary epithelial cells are derived from the embryonic endoderm. Embryological experiments in animal models have demonstrated that liver development occurs through a progressive series of reciprocal tissue interactions between the embryonic endoderm and nearby mesoderm. In the last ten years many of the genes and molecular pathways that regulate hepatogenesis have been identified. Recently application of this knowledge has enabled researchers to produce “hepatic-like” tissue from embryonic stem (ES) cells in vitro, which may ultimately lead to therapeutically useful tissue for transplantation. This review summarizes the current understanding of the molecular pathways controlling liver and biliary system development focusing on studies in the mouse embryo where this process is best understood.
Introduction
The liver is the largest internal organ providing essential metabolic, exocrine and endocrine functions. These include production of bile, metabolism of dietary compounds, detoxification, regulation of glucose levels through glycogen storage and control of blood homeostasis by secretion of clotting factors and serum proteins such as Albumin.
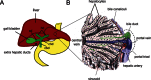
Hepatocytes are the principal cell type in the liver accounting for ∼70% of the mass of the adult organ. Hepatocytes, along with biliary epithelial cells (BECs; also known as cholangyocytes) are derived from the embryonic endoderm, while the stromal cells, stellate cells, kuppfer cells and blood vessels, are of mesodermal origin (see Fig. 1). The use of animal models, such as the mouse, chicken, zebrafish and Xenopus, as well as primary cell cultures has identified many of the genes and molecular pathways regulating embryonic liver development. These studies show that much of hepatogenesis is evolutionarily conserved and occurs through a progressive series of reciprocal tissue interactions between the embryonic endoderms and nearby mesoderm (see Fig. 2; Zaret, 2008; Zhao and Duncan, 2005). The application of this information has recently enabled researchers to produce “hepatic-like” tissue from embryonic stem (ES) cells in vitro, which may ultimately lead to therapeutically useful tissue for transplantation. This review summarizes the current understanding of liver and biliary system development focusing on studies the in mouse embryo where this process is best understood.

Figure 1
Liver cell lineage. The cell lineage steps during hepatic development (red) from uncommitted endoderm to functional adult hepatocytes and biliary epithelium.
Overview of liver development
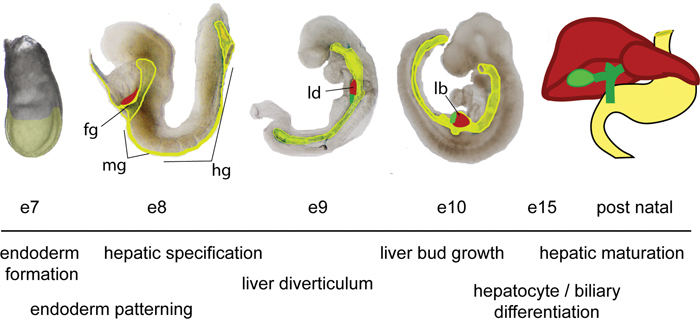
The endoderm germ layer is established during gastrulation and forms a primitive gut tube that is subdivided into foregut, midgut and hindgut regions (see Fig. 2). Fate mapping studies in the mouse embryo at embryonic day 8.0 of gestation (e8.0) indicate that the embryonic liver originates from the ventral foregut endoderm (Tremblay and Zaret, 2005). The first morphological sign of the embryonic liver is the formation of the hepatic diverticulum, an out-pocket of thickened ventral foregut epithelium adjacent to the developing heart at e9.0 (see Fig. 2). The anterior portion of the hepatic diverticulum gives rise to the liver and intrahepatic biliary tree, while the posterior portion forms the gall bladder and extrahepatic bile ducts. At e9.5, the hepatic endoderm cells, known as hepatoblasts delaminate from the epithelium and invade the adjacent septum transversum mesenchyme (STM) to form the liver bud (Houssaint, 1980; Le Douarin, 1975; Medlock and Haar, 1983). The STM contributes fibroblasts and stellate cells of the liver. Between e10–15 the liver bud undergoes a period of accelerated growth as it is vascularized and colonized by hematopoietic cells to become the major fetal hematopoietic organ.
The hepatoblasts are bi-potential and those residing next to the portal veins become BECs that will line the lumen of the intrahepatic bile ducts (IHBD), while the majority of hepatoblasts in the parenchyma differentiate into hepatocytes (Lemaigre, 2003; Shiojiri, 1984). The maturation of functional hepatocytes and the formation of a biliary network connected to the extrahepatic bile ducts (EHBD) are gradual. Beginning at e13 this process continues until after birth to generate the characteristic tissue architecture of the liver.
Cellular architecture of the liver
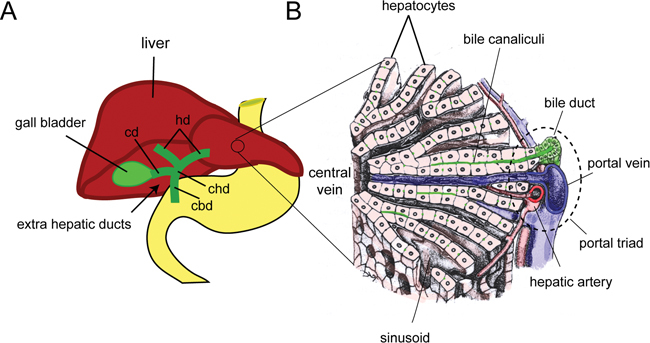
Within the adult liver, the IHBD, portal vein and hepatic artery run in parallel and are referred to as the “portal triad” (see Fig. 3). The portal triad is surrounded by hepatocytes arranged in single cell sheets known as hepatic plates, separated by sinusoid spaces that are connected to a network of blood vessels capillaries. Blood plasma from the portal vein enters the sinusoid space and comes into direct contact with the basal surface of hepatocytes, which absorb metabolites and toxins. Bile is secreted from the apical surface of adjoining hepatocytes into the bile canaliculi (grooves in the cell surface), and then flows though the IHBD to the extrahepatic bile ducts (EHBD), and into the gall bladder where it is stored before release into the duodenum. This cellular architecture is essential for proper hepatic function.
Endoderm formation
In amniotes the definitive endoderm emerges as a sheet of cells from the anterior end of the primitive streak during gastrulation. The multipotent endoderm gives rise to the epithelium of the gastrointestinal and respiratory systems, as well as the glandular and ductal cells of the pancreas, lung, thyroid, thymus and of course the hepatocytes and biliary epithelium. Signaling by the TGFβ growth factor Nodal initiates both endoderm and mesoderm formation in a concentration-dependant manner, with low Nodal doses inducing mesoderm and higher doses inducing endoderm (Shen, 2007; Zorn and Wells, 2007). Nodal signaling stimulates the expression of a core group of endoderm transcription factors including the HMG domain DNA-binding factor Sox17 and the fork head domain proteins Foxa1–3 (HNF3α/β/γ; Zorn and Wells, 2007) which in turn regulate a cascade of genes committing cells to the endoderm lineage.
Endoderm patterning: making the foregut
Initial endoderm patterning is coincident with germ layer formation, as the cells that first emerge from the primitive streak give rise to more anterior endoderm (Lawson and Pedersen, 1987). Anterior endoderm fate requires higher levels of nodal signaling than posterior endoderm cells and expresses Foxa2, which is preferentially required to make anterior definitive endoderm (Dufort et al., 1998; Zorn and Wells, 2007).
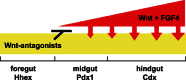
Throughout gastrulation and the early somite stages of development morphogenetic movements turn the endoderm into an epithelial gut tube surrounded by mesoderm. During this time the gut tube epithelium is further patterned along the anterior-posterior (A-P) axis into foregut, midgut and hindgut domains by secreted factors from the adjacent mesoderm. These domains can be identified by the expression of transcription factors such as Hhex in the foregut, Pdx1 in the midgut and Cdx in the posterior endoderm. (Grapin-Botton, 2005; Moore-Scott et al., 2007). The foregut contains the common precursors of the liver, gall bladder, pancreas and lungs (Chalmers and Slack, 2000; Deutsch et al., 2001; Serls et al., 2005; Tremblay and Zaret, 2005). Initially regional identity is plastic and when posterior mesoderm is experimentally recombined with early foregut endoderm it can repress liver and pancreas development, and the endoderm adopts an intestinal fate (Gualdi et al., 1996; Horb and Slack, 2001; Kumar et al., 2003; Okada, 1954; Wells and Melton, 2000).
Foregut development requires repression of Wnt/β-catenin and FGF4
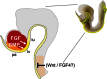
Over lapping temporal and spatial gradients of FGF, Wnt, BMP and retinoic acid secreted from the adjacent mesoderm appear to regulate regional identity of the endoderm (Chen et al., 2004; Dessimoz et al., 2006; Kumar et al., 2003; Martin et al., 2005; McLin et al., 2007; Roberts et al., 1995; Stafford et al., 2004; Tiso et al., 2002). Exactly how these pathways confer regional identity is still unresolved, however recent studies in chick and Xenopus support a model where FGF4 and Wnts secreted from the posterior mesoderm repress foregut fate and promote hindgut development, while Wnt and Fgf4 signaling must be inhibited in the anterior endoderm to establish foregut identity (see Fig. 4; Dessimoz et al., 2006; McLin et al., 2007; Wells and Melton, 2000). Secreted Wnt-antagonists expressed in the foregut endoderm, such as Sfrp5 are predicted to keep Wnt signaling off in the foregut region. Experimentally blocking the activity of the canonical Wnt effector β-catenin in the Xenopus posterior endoderm is sufficient to ectopically activate Hhex expression and result in ectopic liver buds in the intestine (McLin et al., 2007). Amazingly only hours later in development Wnt and FGF signaling have the opposite effects and promote hepatic development.
Hepatic competence
Establishment of the foregut progenitors is important step in hepatogenesis because in vivo only the foregut endoderm (but not the hindgut) is competent to develop into the liver (Fukuda-Taira, 1981; Le Douarin, 1975; Okada, 1954; Okada, 1960; Takata, 1960). This intrinsic hepatic potential is probably due to the expression of transcription factors such as Foxa2, Gata4–6 and Hhex, which have important roles in early foregut organogenesis. In vivo DNA-binding studies in the mouse embryonic endoderm suggest that Foxa and Gata factors bind to enhancer elements of the Albumin gene to enhance chromatin accessibility and increase the ability or “competence” of such hepatic genes to be transcribed (Bossard and Zaret, 1998; Bossard and Zaret, 2000; Gualdi et al., 1996). Consistent with this the conditional deletion of both Foxa1 and Foxa2 from the early foregut prevents liver induction (Lee et al., 2005).
Hepatic induction by FGF and BMP
At early somite stages of development (e8.5; 4–7 somites stage in mice), FGF signals from the developing heart and BMPs from the STM induce hepatic fate in the ventral foregut endoderm (see Fig. 5; Fukuda-Taira, 1981; Gualdi et al., 1996; Jung et al., 1999; Le Douarin, 1964a; Le Douarin, 1964b; Le Douarin, 1975; Rossi et al., 2001). This has been extensively studied using mouse embryo foregut explants, which when isolated at the 2–4-somite stage will express Albumin after 1–2 days in culture if the cardiac mesoderm is present. However, if the cardiac mesoderm is removed, or if either FGF or BMP signaling is blocked, liver induction does not occur (Calmont et al., 2006; Gualdi et al., 1996; Rossi et al., 2001). Furthermore, exogenous FGF1 or FGF2 can replace the cardiac mesoderm and induce Albumin expression in foregut endoderm explants (Jung et al., 1999). BMP signaling however is required, but not sufficient, for hepatic induction in explants and may act in part by maintaining Gata4/6 expression (Rossi et al., 2001).
Analysis of the signal transduction pathways downstream of FGF signaling indicates that the MAP kinase pathway regulates hepatic gene expression whereas the PI3 kinase pathway promotes hepatic growth (Calmont et al., 2006). Gene targeting of FGF and BMP pathway components has not yet revealed a liver-induction phenotype in mice, due to early embryonic lethality and/or functional redundancy. However genetic analysis in zebrafish as well as experiments in chick and Xenopus indicate BMP and FGF signaling cooperatively regulate hepatic liver specification in these species (Chen et al., 2003; Shin et al., 2007; Zhang et al., 2004). In addition, Wnt2bb in the lateral plate mesoderm is required for hepatic specification in zebrafish (Ober et al., 2006). Wnt2 and Wnt2b are also expressed in the mouse lateral plate mesoderm at the time of hepatic induction but their role has yet to be determined.
Mouse explant studies also suggest that different concentrations of FGF are critical for segregating different foregut organ lineages are from a common progenitor cell population, with high, intermediate and low levels of FGF signaling promoting lung, liver and ventral pancreas respectively (Deutsch et al., 2001; Serls et al., 2005). It is still unclear whether it is the proximity, or the length of time that the endoderm is in contact with the cardiac mesoderm that controls the FGF dose. This process may be regulated in part by the homeobox gene Hhex (see Fig. 6), which regulates the proliferation of foregut endoderm cells and causes the epithelium to move in relation to the FGF-secreting cardiac mesoderm during development (Bort et al., 2004).
Liver bud morphogenesis
Shortly after hepatic specification (e8.5 to e9.0) the epithelium begins to express liver genes (Albumin, Afp, Hnf4α) and thickens as the cells transition from a simple cuboidal to a pseudostratified columnar epithelium, thus forming the liver diverticulum (Bort et al., 2006). Between e9.0 to e9.5 the laminin-rich basal layer surrounding the hepatic endoderm breaks down, and the hepatoblasts delaminate and migrate into the STM to form the nascent liver bud (see Fig. 7 and Fig. 8; Bort et al., 2006; Houssaint, 1980; Margagliotti et al., 2008; Medlock and Haar, 1983; Shiojiri and Sugiyama, 2004). Several transcription factors, as well as signals from endothelial cells are required for this process.
Hhex and Gata
Initially expressed throughout the ventral foregut endoderm, the homeodomain factor Hhex becomes enriched in the hepatic endoderm by e8.5 (see Fig. 6) and then persists in all the hepato-biliary cell lineages throughout development (Bogue et al., 2000; Hunter et al., 2007; Thomas et al., 1998). Hhex-/- embryos lack liver, gall bladder and ventral pancreas buds at e10.5 (Keng et al., 2000; Martinez Barbera et al., 2000) and studies indicate that Hhex function is required at multiple time points in the development of these lineages (Bort et al., 2004; Hunter et al., 2007; sections 4, 7 and 8). In addition to controlling foregut endoderm proliferation, Hhex is required for hepatoblast delamination. In e9.0 Hhex-/- embryos the hepatic endoderm transiently expresses liver genes, but the epithelium arrests in a simple columnar state, the hepatoblasts fail to invade the STM (Bort et al., 2006).
The zinc finger transcription factors Gata4 and Gata6 are expressed in a variety of tissues including the foregut endoderm, heart and STM. Mutations in either Gata4 or Gata6 result in early lethality due to defects in the extra embryonic tissue (Duncan, 2005). However in tetraploid chimeric embryos where only the epiblast was derived from Gata4-/- or Gata6-/- ES cells, hepatic development arrested at ∼e9.5 (Watt et al., 2007; Zhao et al., 2005). Similar to the Hhex mutants, hepatic gene expression was initiated but not maintained and the hepatoblasts failed to delaminate. Gata4-/- embryos also lack STM suggesting that part of the defect could be non-cell autonomous (Watt et al., 2007). Compound Gata4-/-; Gata6-/- mutants have early defects in foregut morphogenesis but the liver phenotype has not yet been reported in these embryos (Zhao et al., 2008). The proposed role for Gata factors in hepatic competence (section 3.2) predicts that liver development should not initiate in the compound mutants, which is consistent with the antisense depletion of Gata4, Gata5 and Gata6 in zebrafish and Xenopus embryos (Afouda et al., 2005; Holtzinger and Evans, 2005; Shin et al., 2007).
Prox1, Onecut and the ECM
The homeodomain transcription factors Prox1, Onecut-1 (OC-1, also known as Hnf6) and Onecut-2 (OC-2) also regulate hepatoblast delamination, but act slightly later than Hhex and Gata. Prox1 is expressed in the hepatic epithelium by e8.5 (Burke and Oliver, 2002). In Prox1-/- embryos hepatoblasts are specified and begin to proliferate, but the basal lamina fails to degrade and the cells remain trapped in the hepatic diverticulum (Sosa-Pineda et al., 2000). OC-1 and OC-2 are expressed in the foregut endoderm and hepatoblasts, and are redundantly required for the timely degradation of the basal lamina (Margagliotti et al., 2007). At e9.5, OC-1;OC-2 double mutants resemble the Prox1-/- phenotype, but later hepatoblast invasion recovers resulting in a hypoplastic fetal liver. These mutant livers also have bile duct and gall bladder defects due to later roles of OC-1 and OC-2 (sections 7.2 & 8).
Hepatoblasts normally down regulate E-cadherin as they migrate into the STM, however this does not occur properly in Prox1 and OC1;OC2 mutants. Studies suggest that Prox1 and Onecut factors control hepatoblast migration by regulating the expression of extra cellular matrix (ECM) proteins and ECM remodeling enzymes such as matrix metalloproteinases (MMPs; Margagliotti et al., 2007; Medico et al., 2001; Papoutsi et al., 2007). Hepatoblasts and the STM express several MMPs at e9.5 and pharmacological inhibition of MMP activity suppresses hepatoblast migration in culture (Margagliotti et al., 2008). The importance of cell-ECM interactions is also illustrated by the fact that hepatoblasts deficient for the laminin receptor β1-integrin are unable to colonize the liver bud (Fassler and Meyer, 1995). Small GTPases, well known for regulating cell migration, also appear to be involved because hepatoblasts fail to invade the STM in embryos with null mutations in the Pccmt gene, which encodes a GTPase modifying enzyme (Lin et al., 2002).
Endothelial signals
At e9.0, prior to vascularization of the liver bud, endothelial precursor cells lay between the hepatic epithelium and the STM (see Fig. 7). This close contact with blood vessels persists as hepatoblasts migrate into the stroma. Null mutations in the vascular endothelial growth factor receptor gene Vegfr-2 (also known as Flk-1) results in embryos that lack endothelial cells and the hepatoblasts fail to delaminate in these embryos (Matsumoto et al., 2001). In addition angiogenesis inhibitors repress liver bud growth in culture, suggesting that endothelial cells provide unknown paracrine factors promoting hepatoblast migration and/or proliferation. One candidate that has emerged from chick embryo studies is the Glial-derived neurotrophic factor, Neurturin, which is secreted from blood vessels and acts as a hepatoblast chemoattractant via the GFRα2 receptor (Tatsumi et al., 2007).
Liver bud growth
Between e9.5 to e15 the liver bud undergoes tremendous growth and becomes the major site of fetal haematopoiesis. This growth is regulated by paracrine signals from hepatic mesenchyme, as well as by genes that act intrinsically in the hepatoblasts. Mutations in many of these genes result in a similar embryonic lethality between e10-e16 due to impaired hepatoblast proliferation and/or increased cell death, which often causes severe anemia because the defective liver cannot support fetal haematopoiesis.
Mesenchymal signals
The STM and hepatic mesenchyme (e.g. stellate cells) secrete a variety of growth factors including; FGF, BMP, HGF, Wnt, TGFβ and RA that promote hepatoblast migration, proliferation, and survival (see Fig. 9).
In addition to their earlier role in hepatic specification, FGF (via PI3 kinase) and BMP signaling also promote liver bud growth (Berg et al., 2007; Calmont et al., 2006; Jung et al., 1999; Rossi et al., 2001; Sekhon et al., 2004; Shin et al., 2007; Yanai et al., 2008). Hepatocyte Growth Factor (HGF) signaling through its tyrosine kinase receptor c-Met (Defrances et al., 1992; Iida et al., 2003; Ishikawa et al., 2001) is also required for hepatoblast proliferation (Birchmeier et al., 2003; Bladt et al., 1995; Moumen et al., 2007; Sachs et al., 2000; Schmidt et al., 1995). In addition to being a hepatocyte mitogen HGF also promotes hepatoblast migration (Block et al., 1996; Medico et al., 2001; Michalopoulos et al., 1993), in part by activating the small GTPase Arf6 (Suzuki et al., 2006).
Although Wnt/β-catenin signaling appears to repress liver fate during earlier endoderm patterning stages of development (section 3.1), by e10 in the liver bud β-catenin has the opposite effect and promotes hepatic growth (McLin et al., 2007; Micsenyi et al., 2004; Monga et al., 2003; Suksaweang et al., 2004; Tan et al., 2008). While the Wnt ligands involved are unclear (Zeng et al., 2007), in the chick Wnt9a is expressed in hepatic fibroblasts, and antisense depletion of Wnt9a inhibits liver bud growth in culture (Matsumoto et al., 2008). Multiple TGFβ ligands are also expressed in the liver bud mesenchyme (Pelton et al., 1991). Embryos with compound heterozygous mutations in the TGFβ transcriptional mediators Smad2 and Smad3 have hypoplastic livers (Weinstein et al., 2001), as do embryos deficient for the Smad interacting protein, Elf5 (Tang et al., 2003) and the receptor TGFβRIII (Stenvers et al., 2003)
Liver bud growth also requires retinoic acid signaling (Wang et al., 2006), which is controlled in part by the zinc finger transcription factor WT1 expressed in the STM and stellate cells (Ijpenberg et al., 2007). The homeobox genes Hlx and Lhx2 as well as N-myc are also expressed in the STM, and null mutations in each of these results in reduced hepatoblast proliferation and increased apoptosis (Giroux and Charron, 1998; Hentsch et al., 1996; Porter et al., 1997; Wandzioch et al., 2004), suggesting that they regulate the production of paracrine signals from the mesenchyme. Exactly how these different transcription factors and signaling pathways coordinate hepatic growth remains to be determined, but extensive cross talk appears to exist (Apte et al., 2006; Rossi et al., 2001; Weinstein et al., 2001). For example, explant cultures suggest that HGF and TGFβ signaling act in parallel converging on β1-integrin regulation (Weinstein et al., 2001). Moreover, FGF and HGF signaling stimulate many of the same intracellular kinase cascades (e.g. MAPK, JNK, Pi3K), and both have been reported to stimulate the activity of β-catenin in the liver bud, suggesting crosstalk with the Wnt pathway (Berg et al., 2007; Monga et al., 2002; Sekhon et al., 2004).
Hepatoblast proliferation and survival
A number of genes encoding regulators of proliferation, cell survival or metabolic stress are also required for liver bud growth including; c-jun (Eferl et al., 1999; Hilberg et al., 1993), Xbp1 (Reimold et al., 2000), Jumonji (Motoyama et al., 1997), Foxm1b (Krupczak-Hollis et al., 2004), MTF-1 (Gunes et al., 1998), Nrf1 (Chen et al., 2003), Tbx3 (Suzuki et al., 2008) and signal transduction components K-ras (Johnson et al., 1997), Pi3kr1 (Fruman et al., 2000), Raf1 (Mikula et al., 2001) and Sek1 (Ganiatsas et al., 1998; Nishina et al., 1999; Watanabe et al., 2002). Regulation of the inflammatory cytokine Tumor Necrosis Factor alpha (TNFα) is also important and embryos lacking components of the anti-apoptotic NFκβ complex exhibit TNFα mediated hepatoblast apoptosis (Beg et al., 1995; Bonnard et al., 2000; Doi et al., 1999; Li et al., 1999; Rosenfeld et al., 2000).
Differentiation of hepatocytes and biliary epithelial cells
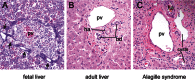
The differentiation of bi-potential hepatoblasts into hepatocytes or BECs begins around e13 of mouse development. Initially hepatoblasts express genes associated with both adult hepatocytes (Hnf4α, Albumin) and BECs (cytokeratin-19), as well as fetal liver genes such as α-fetoprotein (Afp). Hepatoblasts in contact with the portal vein form a monolayer, and then a bi-layer, of cuboidal biliary precursors that increase cytokeratin-19 (CK-19) expression and down-regulate hepatic genes. Between e17 and into the perinatal period focal dilations appear in the bi-layer and these become surrounded by portal mesenchyme to form IHBD, while the remaining bi-layer cells regress (see Fig. 10). This process, which involves tubulogenesis and apoptosis, is known as ductal plate remodeling. (Lemaigre, 2003; Sergi et al., 2000). Hepatoblasts in the liver parenchyma that are not in contact with portal veins gradually differentiate into mature hepatocytes. At e17 hepatocytes acquire their characteristic epithelial morphology arranged in hepatic chords with bile canaliculi on the apical surfaces. While defects in early liver bud growth are often embryonic lethal, disruption in hepatocyte maturation/function or ductal plate remodeling malformations (see Fig. 11) are observed in many human disorders (Sergi et al., 2000; Suchy, 2003; Desmet, 2005).
Hepatocyte differentiation
During mid-gestation the haematopoietic cells in the liver secrete the cytokine Oncostatin M (OSM), which in combination with glucocorticoid hormones, HGF and Wnt promotes hepatocyte differentiation (Kamiya et al., 2001; Matsui et al., 2002; Michalopoulos et al., 2003; Suzuki et al., 2003; Tan et al., 2008). OSM induces metabolic maturation by activating the gp130 receptor and a JAK/Stat3 signaling pathway (Ito et al., 2000; Kamiya et al., 1999), while promoting morphological maturation into polarized epithelium via K-ras and E-cadherin (Imamura et al., 2007; Matsui et al., 2002). Some evidence suggests that HGF and OSM activity is balanced by TNFα, which inhibits maturation and maintains the proliferative capacity of fetal hepatocytes, allowing the liver to grow to the appropriate size before differentiating (Kamiya and Gonzalez, 2004).
These secreted factors act in part by regulating a number of liver-enriched transcription factors including C/EBPα, HNF1α, Foxa1–3, nuclear hormone receptors and HNF4α (see Fig. 12), which function in a complex inter-regulatory network to control hepatocyte gene expression (Cheng et al., 2006; Kyrmizi et al., 2006; Odom et al., 2004; Qu et al., 2007). Genetic analysis has confirmed a role for HNF4α, C/EBPα and HNF1α in hepatocyte differentiation. HNF4α is first expressed in hepatoblasts at e9.0 and Hnf4α-/- fetal hepatocytes fail to express many mature hepatic enzymes and lack normal hepatocyte morphology (Li et al., 2000; Parviz et al., 2003; Watt et al., 2003). Genome scale chromatin immunoprecipitation assays suggest that HNF4α binds to the promoters of nearly half of the genes expressed in the mouse liver (Odom et al., 2004), including genes encoding cell adhesion and junctional proteins (Battle et al., 2006) important in hepatocyte epithelial structure (Konopka et al., 2007; Satohisa et al., 2005). C/ebpα-/- and Hnf1α-/- mice die neonatally from hypoglycemia caused by impaired hepatocyte maturation and defective glycogen storage (Pontoglio et al., 1996; Wang et al., 1995), similar to mutations in the OSM receptor gp130 (Kamiya et al., 1999). C/EBPα appears to inhibit hepatoblast proliferation and biliary development, while mediating HGF-induced hepatic gene expression (Rastegar et al., 2000; Shiojiri et al., 2004; Soriano et al., 1995; Suzuki et al., 2003; Tomizawa et al., 1998; Yamasaki et al., 2006)
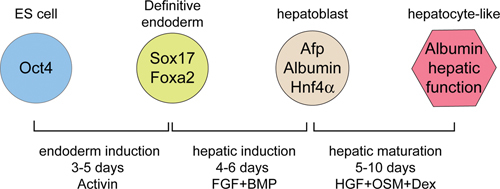
Figure 13In vitro hepatic differentiation from ES cells.
The diagram depicts a generalized protocol summarized from the work of several labs that have applied developmental paradigms to mouse and human ES cells to direct the differentiation of hepatic-like cell in vitro. The factors added to the cultures, the durations of exposure, and the developmental step that these treatments are meant to mimic are indicated below. The cell types and key lineage specific marker genes expressed in those cells are indicated.
Biliary epithelial cells
TGFβ, Wnt and Notch are candidate signals from periportal mesenchyme that promotes BEC development (see Fig. 12). There is evidence that a TGFβ signaling gradient emanating from the portal region promotes biliary differentiation in the adjacent hepatoblasts (Clotman et al., 2005; Clotman and Lemaigre, 2006; Weinstein et al., 2001). Wnt/β-catenin signaling also promotes BEC development (Decaens et al., 2008; Hussain et al., 2004; Monga et al., 2003; Tan et al., 2008), and may act in part by stimulating the expression of EGF, which along with HGF can induce the formation of biliary structures in cultured hepatocytes (Block et al., 1996; Michalopoulos et al., 2003; Tan et al., 2005).
Human Alagille syndrome (OMIN #118450) patients with autosomal dominant mutations in the Notch ligand gene Jagged-1 (Li et al., 1997; Oda et al., 1997; Yuan et al., 2006) have a paucity of IHBD (see Fig. 11), as do mice with compound heterozygous Jagged-1; Notch-2 mutations (McCright et al., 2002). Recent analyses suggest that Jagged-1 expressed in the portal mesenchyme activates Notch-2 in adjacent hepatoblast, and this is required to maintain (but not initiate) BEC differentiation and for proper bile duct morphogenesis (Kodama et al., 2004; Loomes et al., 2007; Lozier et al., 2008; Tanimizu and Miyajima, 2004).
In response to the mesenchyme signals the periportal hepatoblasts down regulate pro-hepatic factors HNF4α, Tbx3 and C/EBP (which repress BEC development) and increase expression of BEC transcription factors OC1/HNF6, OC2 and HNF1β (see Fig. 12; Rastegar et al., 2000; Shiojiri et al., 2004; Suzuki et al., 2008; Tanimizu and Miyajima, 2004; Yamasaki et al., 2006). Fetal livers deficient for OC1/HNF6, OC2, HNF1β or Hhex exhibit excessive biliary cysts that fail to form proper IHBDs (Clotman et al., 2005; Clotman et al., 2002; Coffinier et al., 2002; Hunter et al., 2007). At e13-e15 Hhex, OC1 and OC2 appear to regulate early hepato-biliary lineage segregation and attenuate premature BEC differentiation. OC1 and OC2 may act in part by modulating the response of hepatoblasts to TGFβ signaling (Clotman et al., 2005). Later in development Hhex and OC1 function upstream of HNF1β and are required for ductal plate remodeling (Clotman et al., 2002; Coffinier et al., 2002; Hunter et al., 2007; Matthews et al., 2004).
In addition to reduced hepatoblast proliferation Foxm1b-/- fetal livers lack IHBDs (Krupczak-Hollis et al., 2004). It is currently unclear how Foxm1b fits into the genetic pathway of BEC development. Interestingly cell lines have been derived from bipotential fetal hepatoblasts that can be maintained and differentiated in vitro (Rogler, 1997; Strick-Marchand and Weiss, 2002). These are likely to be instrumental in dissecting the genetic cascades regulating BEC/hepatocyte lineage segregation as well as provide potential source of hepatic tissue for tissue replacement therapy (section 9; Ader et al., 2006; Strick-Marchand et al., 2004; Strick-Marchand and Weiss, 2003).
Gall bladder and extrahepatic bile ducts
Although BECs line the lumen of both the intrahepatic and extrahepatic biliary tract they have distinct developmental origins (Roskams and Desmet, 2008), with the BECs of the gall bladder and extrahepatic bile ducts (EHBD) arising from the caudal portion of the hepatic diverticulum. While some of the same genes control BEC fate in both the IHBD and EHBD, other aspects of their development appears distinct. For example HNF6, HNF1b, Hhex and the Notch signaling effector Hex1 are required for IHBD as well as for gall bladder and EHBD development (Clotman et al., 2002; Coffinier et al., 2002; Hunter et al., 2007). However haploinsufficency of the transcription factor Foxf1 results in a lack of EHBDs and a small gall bladder, but IHBDs are unaffected (Kalinichenko et al., 2002). Foxf1 is expressed in the STM suggesting that it acts indirectly on the hepato-biliary epithelium. Evidence for such an extra-cellular signal has come from zebrafish where Fgf10 in the lateral plate mesoderm regulates the boundary between the liver, gall bladder, ventral pancreas and EHBD (Dong et al., 2007).
Liver stem cells
The adult liver has a remarkable regenerative capacity and can completely re-grow when up to 70% of its mass is removed (Fausto et al., 2006; Michalopoulos, 2007). However this ability is impaired in numerous diseases such as advanced cirrhosis and hepatitis resulting life threatening liver failure for which organ transplantation is currently the only clinical option. The implantation of isolated hepatocytes or the use of bio-artificial liver devices to provide limited liver function, are also becoming potential options (Dan and Yeoh, 2008; Horslen and Fox, 2004; McKenzie et al., 2008). However, the scarcity of organ donors and the difficulty in culturing adult hepatocytes are serious limitation to this approach. Hepatic tissue made from stem cells holds the promise of an unlimited source of material for transplantation. Moreover the availability of large quantities of human hepatic tissue would facilitate the design of new pharmaceuticals as testing for liver toxicity is a common step in drug development. Researchers have attempted to generate hepatocytes from a variety of adult, fetal and embryonic stem cell sources (Dan and Yeoh, 2008; Lavon and Benvenisty, 2005; Oertel and Shafritz, 2008). These efforts, particularly the recent successes with embryonic stem (ES) cells, have been greatly facilitated by our understanding of embryonic liver development.
Regeneration and adult hepatic stem cells
The liver regenerates primarily by the proliferation of mature hepatocytes (Fausto et al., 2006; Michalopoulos, 2007). However, the adult liver also contains hepatic progenitor cells that are activated when hepatocyte proliferation is inhibited, such as in severe cirrhosis (Fausto, 2004; Oertel and Shafritz, 2008). The hepatic progenitors appear to reside in the small terminal bile ducts and when activated they proliferate giving rise to a cell population called “oval cells”, which can differentiate into both hepatocytes and BECs (Fougere-Deschatrette et al., 2006; Oertel and Shafritz, 2008). The bipotential nature of oval cells suggests that they originate from fetal hepatoblasts that remain undifferentiated in a stem cell niche within the ducts (Schmelzer et al., 2006). Rodent Oval cell lines have been generated (Fougere-Deschatrette et al., 2006; Oertel and Shafritz, 2008) and provide useful experimental models, but the exact origin and characteristics of hepatic progenitors in vivo are still obscure (Dan and Yeoh, 2008; Jelnes et al., 2007; Ochsner et al., 2007; Oertel et al., 2008; Yovchev et al., 2008). Interestingly, many of the developmental pathways that regulate hepatogenesis in the embryo, such as HGF, FGF, OSM, TNFα and Wnt appear to control Oval cell activation (Apte et al., 2008; Bird et al., 2008; Hu et al., 2007) as well as hepatocyte proliferation during regeneration. (Fausto, 2004; Kelley-Loughnane et al., 2002; Michalopoulos, 2007; Tan et al., 2006).
Making liver from ES cells
ES cells are capable of unlimited self-renewal and can differentiate into any cell type in the adult. For years investigators attempted to promote hepatic differentiation in ES cells, however initial results were hampered by variability, low yields and the heterogeneous nature of the cultures (Lavon and Benvenisty, 2005). The breakthrough came from recapitulating the pathways controlling normal hepatic development in ES cells.
The critical first step was the demonstration that mouse and human ES cell could efficiently generate definitive endoderm (DE) tissue (∼80% of the cells) by treating the cultures with high concentrations of the TGFβ family ligand Activin, to mimic the role of Nodal in the gastrula embryo (D’Amour et al., 2005; Kubo et al., 2004; Yasunaga et al., 2005). Remarkably Activin treated ES cells express mesendoderm genes in a temporal fashion nearly identical to mammalian gastrulation. After a few days most of the cells express definitive endoderm genes (Sox17 and Foxa2), while mesoderm, pluripotency (e.g. Oct4) and extra embryonic endoderm genes are down regulated (D’Amour et al., 2005; Kubo et al., 2004). Using this DE tissue as a starting material, a number of groups have generated hepatic-like cells from mouse and human ES cells in culture (reviewed in Dan and Yeoh, 2008; Oertel and Shafritz, 2008). While the details of the 15–20 day long differentiation protocols vary, most involve plating the DE on matrix (e.g. collagen) to mimic the hepatic ECM and then added FGF and/or BMP to mimic hepatic induction. This is followed by some combination of HGF, OSM, FGF and Dexamethasone to expand the hepatoblast population and to promote hepatic maturation (Agarwal et al., 2008; Baharvand et al., 2007; Cai et al., 2007; Chen et al., 2006; Cho et al., 2008; Gouon-Evans et al., 2006; Hay et al., 2008; Soto-Gutierrez et al., 2007; Teratani et al., 2005). A detailed understanding of the stage specific genes expressed during normal liver development has allowed researchers to monitor the progress of hepatic differentiation in vitro.
Using these developemental protocols on mouse and human ES cells, researchers have successfully generated cultures where up to 70% of the cells exhibit a “hepatocyte-like” phenotype. These cells exhibit many features of hepatocytes including; (1) expression of hepatic enzymes, (2) hepatocyte morphology, (3) robust glycogen storage, (4) uptake and metabolizism of drugs and (5) secretion of albumin. Finally several groups have found that transfusion of ES derived hepatocytes into mice with various liver injury models exhibited a modest rescue of liver function, although engraftment of the cells into the host liver was very low (Agarwal et al., 2008; Cai et al., 2007; Gouon-Evans et al., 2006; Heo et al., 2006; Sharma et al., 2008). ES cell derived hepatocytes implanted in a bio-artificial liver devices have also been shown to partially reverse fulminant liver failure and 90% hepatectomy in mice (Cho et al., 2008; Soto-Gutierrez et al., 2006).
While these results provide a promising proof of principle there are still limitations. In addition to poor engraftment, the hepatic-like cells produced still have characteristics of fetal hepatocytes such as Afp expression (Agarwal et al., 2008; Cai et al., 2007; Gouon-Evans et al., 2006), suggesting that the cells are not fully differentiated. It is likely that continued investigations into normal liver development would enhance our ability to differentiate ES cells into therapeutically useful hepatic tissue in vitro.
Conclusions
In the last ten years researchers have uncovered many of the genes regulating hepatogenesis. One of the emerging themes is that many of the same signaling pathways and transcription factors are used reiteratively, and have different functions at different times in development. A major challenge now is to understand how all these genes and pathways interact in a precise temporal sequence to direct a naïve endoderm cell into a functional hepatocyte. In addition there are still gaps in our knowledge of both early foregut patterning and later aspects of hepatic maturation. The recent rise of alternative model organisms such as genetic screens in fish and functional assays in Xenopus are helping to accelerate our understanding of early hepatogenesis (Cheng et al., 2006; Ober et al., 2006; Sadler et al., 2005). Finally the use of hepatic progenitor cell cultures as well as in vitro liver development from ES cells will be valuable models for studying hepatic differentiation.
Acknowledgement
I apologize to authors whose work I could not describe due to space constraints. I am grateful to Gail Deutsch, Jay Kormish, Chris Mayhew, Scott Rankin, Jason Spence and Jim Wells for comments on the manuscript and for providing images for the figures. Liver research in the Zorn lab is supported by the NIH grant DK070858.
References
- Ader T, Norel R, Levoci L, Rogler L.E. Transcriptional profiling implicates TGFbeta/BMP and Notch signaling pathways in ductular differentiation of fetal murine hepatoblasts. Mech Dev. 2006;123:177–194. [PubMed: 16412614]
- Afouda B.A, Ciau-Uitz A, Patient R. GATA4, 5 and 6 mediate TGFbeta maintenance of endodermal gene expression in Xenopus embryos. Development. 2005;132:763–774. [PubMed: 15659482]
- Agarwal S, Holton K.L, Lanza R. Efficient Differentiation of Functional Hepatocytes from Human Embryonic Stem Cells. Stem Cells. 2008 [PubMed: 18292207]
- Apte U, Thompson M.D, Cui S, Liu B, Cieply B, Monga S.P. Wnt/beta-catenin signaling mediates oval cell response in rodents. Hepatology. 2008;47:288–295. [PubMed: 17929301]
- Apte U, Zeng G, Muller P, Tan X, Micsenyi A, Cieply B, Dai C, Liu Y, Kaestner K.H, Monga S.P. Activation of Wnt/beta-catenin pathway during hepatocyte growth factor-induced hepatomegaly in mice. Hepatology. 2006;44:992–1002. [PubMed: 17006939]
- Baharvand H, Hashemi S.M, Shahsavani M. Differentiation of human embryonic stem cells into functional hepatocyte-like cells in a serum-free adherent culture condition. Differentiation. 2007 [PubMed: 18177418]
- Battle M.A, Konopka G, Parviz F, Gaggl A.L, Yang C, Sladek F.M, Duncan S.A. Hepatocyte nuclear factor 4alpha orchestrates expression of cell adhesion proteins during the epithelial transformation of the developing liver. Proc Natl Acad Sci USA. 2006;103:8419–8424. [PMC free article: PMC1482507] [PubMed: 16714383]
- Beg A.A, Sha W.C, Bronson R.T, Ghosh S, Baltimore D. Embryonic lethality and liver degeneration in mice lacking the RelA component of NF-kappa B. Nature. 1995;376:167–170. [PubMed: 7603567]
- Berg T, Rountree C.B, Lee L, Estrada J, Sala F.G, Choe A, Veltmaat J.M, De Langhe S, Lee R, Tsukamoto H. et al. Fibroblast growth factor 10 is critical for liver growth during embryogenesis and controls hepatoblast survival via beta-catenin activation. Hepatology. 2007;46:1187–1197. [PMC free article: PMC3494299] [PubMed: 17668871]
- Birchmeier C, Birchmeier W, Gherardi E, Vande Woude G.F. Met, metastasis, motility and more. Nat Rev Mol Cell Biol. 2003;4:915–925. [PubMed: 14685170]
- Bird T.G, Lorenzini S, Forbes S.J. Activation of stem cells in hepatic diseases. Cell Tissue Res. 2008;331:283–300. [PMC free article: PMC3034134] [PubMed: 18046579]
- Bladt F, Riethmacher D, Isenmann S, Aguzzi A, Birchmeier C. Essential role for the c-met receptor in the migration of myogenic precursor cells into the limb bud. Nature. 1995;376:768–771. [PubMed: 7651534]
- Block G.D, Locker J, Bowen W.C, Petersen B.E, Katyal S, Strom S.C, Riley T, Howard T.A, Michalopoulos G.K. Population expansion, clonal growth, and specific differentiation patterns in primary cultures of hepatocytes induced by HGF/SF, EGF and TGF alpha in a chemically defined (HGM) medium. J Cell Biol. 1996;132:1133–1149. [PMC free article: PMC2120765] [PubMed: 8601590]
- Bogue C.W, Ganea G.R, Sturm E, Ianucci R, Jacobs H.C. Hex expression suggests a role in the development and function of organs derived from foregut endoderm. Dev Dyn. 2000;219:84–89. [PubMed: 10974674]
- Bonnard M, Mirtsos C, Suzuki S, Graham K, Huang J, Ng M, Itie A, Wakeham A, Shahinian A, Henzel W.J. et al. Deficiency of T2K leads to apoptotic liver degeneration and impaired NF-kappaB-dependent gene transcription. Embo J. 2000;19:4976–4985. [PMC free article: PMC314216] [PubMed: 10990461]
- Bort R, Martinez-Barbera J.P, Beddington R.S, Zaret K.S. Hex homeobox gene-dependent tissue positioning is required for organogenesis of the ventral pancreas. Development. 2004;131:797–806. [PubMed: 14736744]
- Bort R, Signore M, Tremblay K, Barbera J.P, Zaret K.S. Hex homeobox gene controls the transition of the endoderm to a pseudostratified, cell emergent epithelium for liver bud development. Dev Biol. 2006;290:44–56. [PubMed: 16364283]
- Bossard P, Zaret K.S. GATA transcription factors as potentiators of gut endoderm differentiation. Development. 1998;125:4909–4917. [PubMed: 9811575]
- Bossard P, Zaret K.S. Repressive and restrictive mesodermal interactions with gut endoderm: possible relation to Meckel's Diverticulum. Development. 2000;127:4915–4923. [PubMed: 11044405]
- Burke Z, Oliver G. Prox1 is an early specific marker for the developing liver and pancreas in the mammalian foregut endoderm. Mech Dev. 2002;118:147–155. [PubMed: 12351178]
- Cai J, Zhao Y, Liu Y, Ye F, Song Z, Qin H, Meng S, Chen Y, Zhou R, Song X. et al. Directed differentiation of human embryonic stem cells into functional hepatic cells. Hepatology. 2007;45:1229–1239. [PubMed: 17464996]
- Calmont A, Wandzioch E, Tremblay K.D, Minowada G, Kaestner K.H, Martin G.R, Zaret K.S. An FGF response pathway that mediates hepatic gene induction in embryonic endoderm cells. Dev Cell. 2006;11:339–348. [PubMed: 16950125]
- Chalmers A.D, Slack J.M. The Xenopus tadpole gut: fate maps and morphogenetic movements. Development. 2000;127:381–392. [PubMed: 10603354]
- Chen L, Kwong M, Lu R, Ginzinger D, Lee C, Leung L, Chan J.Y. Nrf1 is critical for redox balance and survival of liver cells during development. Mol Cell Biol. 2003;23:4673–4686. [PMC free article: PMC164851] [PubMed: 12808106]
- Chen Y, Jurgens K, Hollemann T, Claussen M, Ramadori G, Pieler T. Cell-autonomous and signal-dependent expression of liver and intestine marker genes in pluripotent precursor cells from Xenopus embryos. Mech Dev. 2003;120:277–288. [PubMed: 12591597]
- Chen Y, Pan F.C, Brandes N, Afelik S, Solter M, Pieler T. Retinoic acid signaling is essential for pancreas development and promotes endocrine at the expense of exocrine cell differentiation in Xenopus. Dev Biol. 2004;271:144–160. [PubMed: 15196957]
- Chen Y, Soto-Gutierrez A, Navarro-Alvarez N, Rivas-Carrillo J.D, Yamatsuji T, Shirakawa Y, Tanaka N, Basma H, Fox I.J, Kobayashi N. Instant hepatic differentiation of human embryonic stem cells using activin A and a deleted variant of HGF. Cell Transplant. 2006;15:865–871. [PubMed: 17299990]
- Cheng W, Guo L, Zhang Z, Soo H.M, Wen C, Wu W, Peng J. HNF factors form a network to regulate liver-enriched genes in zebrafish. Dev Biol. 2006;294:482–496. [PubMed: 16631158]
- Cho C.H, Parashurama N, Park E.Y, Suganuma K, Nahmias Y, Park J, Tilles A.W, Berthiaume F, Yarmush M.L. Homogeneous differentiation of hepatocyte-like cells from embryonic stem cells: applications for the treatment of liver failure. Faseb J. 2008;22:898–909. [PubMed: 17942827]
- Clotman F, Jacquemin P, Plumb-Rudewiez N, Pierreux C.E, Van der Smissen P, Dietz H.C, Courtoy P.J, Rousseau G.G, Lemaigre F.P. Control of liver cell fate decision by a gradient of TGF beta signaling modulated by Onecut transcription factors. Genes Dev. 2005;19:1849–1854. [PMC free article: PMC1186184] [PubMed: 16103213]
- Clotman F, Lannoy V.J, Reber M, Cereghini S, Cassiman D, Jacquemin P, Roskams T, Rousseau G.G, Lemaigre F.P. The onecut transcription factor HNF6 is required for normal development of the biliary tract. Development. 2002;129:1819–1828. [PubMed: 11934848]
- Clotman F, Lemaigre F. P. Control of hepatic differentiation by activin/TGFbeta signaling. Cell Cycle. 2006;5:168–171. [PubMed: 16357531]
- Coffinier C, Gresh L, Fiette L, Tronche F, Schutz G, Babinet C, Pontoglio M, Yaniv M, Barra J. Bile system morphogenesis defects and liver dysfunction upon targeted deletion of HNF1beta. Development. 2002;129:1829–1838. [PubMed: 11934849]
- D’Amour K.A, Agulnick A.D, Eliazer S, Kelly O.G, Kroon E, Baetge E.E. Efficient differentiation of human embryonic stem cells to definitive endoderm. Nat Biotechnol. 2005;23:1534–1541. [PubMed: 16258519]
- Dan Y.Y, Yeoh G.C. Liver stem cells: a scientific and clinical perspective. J Gastroenterol Hepatol. 2008;23:687–698. [PubMed: 18410603]
- Decaens T, Godard C, de Reynies A, Rickman D.S, Tronche F, Couty J.P, Perret C, Colnot S. Stabilization of beta-catenin affects mouse embryonic liver growth and hepatoblast fate. Hepatology. 2008;47:247–258. [PubMed: 18038450]
- Defrances M.C, Wolf H.K, Michalopoulos G.K, Zarnegar R. The presence of hepatocyte growth factor in the developing rat. Development. 1992;116:387–395. [PubMed: 1286614]
- Desmet V.J. Cystic diseases of the liver. From embryology to malformations. Gastroenterol Clin Biol. 2005;29:858–860. [PubMed: 16294158]
- Dessimoz J, Opoka R, Kordich J.J, Grapin-Botton A, Wells J.M. FGF signaling is necessary for establishing gut tube domains along the anterior-posterior axis in vivo. Mech Dev. 2006;123:42–55. [PubMed: 16326079]
- Deutsch G, Jung J, Zheng M, Lora J, Zaret K.S. A bipotential precursor population for pancreas and liver within the embryonic endoderm. Development. 2001;128:871–881. [PubMed: 11222142]
- Doi T.S, Marino M.W, Takahashi T, Yoshida T, Sakakura T, Old L. J, Obata Y. Absence of tumor necrosis factor rescues RelA-deficient mice from embryonic lethality. Proc Natl Acad Sci USA. 1999;96:2994–2999. [PMC free article: PMC15883] [PubMed: 10077625]
- Dong P.D, Munson C.A, Norton W, Crosnier C, Pan X, Gong Z, Neumann C.J, Stainier D.Y. Fgf10 regulates hepatopancreatic ductal system patterning and differentiation. Nat Genet. 2007;39:397–402. [PubMed: 17259985]
- Dufort D, Schwartz L, Harpal K, Rossant J. The transcription factor HNF3beta is required in visceral endoderm for normal primitive streak morphogenesis. Development. 1998;125:3015–3025. [PubMed: 9671576]
- Duncan S.A. Generation of embryos directly from embryonic stem cells by tetraploid embryo complementation reveals a role for GATA factors in organogenesis. Biochem Soc Trans. 2005;33:1534–1536. [PubMed: 16246163]
- Eferl R, Sibilia M, Hilberg F, Fuchsbichler A, Kufferath I, Guertl B, Zenz R, Wagner E.F, Zatloukal K. Functions of c-Jun in liver and heart development. J Cell Biol. 1999;145:1049–1061. [PMC free article: PMC2133137] [PubMed: 10352021]
- Fassler R, Meyer M. Consequences of lack of beta 1 integrin gene expression in mice. Genes Dev. 1995;9:1896–1908. [PubMed: 7544313]
- Fausto N. Liver regeneration and repair: hepatocytes, progenitor cells, and stem cells. Hepatology. 2004;39:1477–1487. [PubMed: 15185286]
- Fausto N, Campbell J.S, Riehle K.J. Liver regeneration. Hepatology. 2006;43:S45–53. [PubMed: 16447274]
- Fougere-Deschatrette C, Imaizumi-Scherrer T, Strick-Marchand H, Morosan S, Charneau P, Kremsdorf D, Faust D.M, Weiss M.C. Plasticity of hepatic cell differentiation: bipotential adult mouse liver clonal cell lines competent to differentiate in vitro and in vivo. Stem Cells. 2006;24:2098–2109. [PubMed: 16946000]
- Fruman D.A, Mauvais-Jarvis F, Pollard D.A, Yballe C.M, Brazil D, Bronson R.T, Kahn C.R, Cantley L.C. Hypoglycaemia, liver necrosis and perinatal death in mice lacking all isoforms of phosphoinositide 3-kinase p85 alpha. Nat Genet. 2000;26:379–382. [PubMed: 11062485]
- Fukuda-Taira S. Hepatic induction in the avian embryo: Specificity of reactive endoderm and inductive mesoderm. J Embryol exp Morph. 1981;63:111–125. [PubMed: 7310284]
- Ganiatsas S, Kwee L, Fujiwara Y, Perkins A, Ikeda T, Labow M. A, Zon L. I. SEK1 deficiency reveals mitogen-activated protein kinase cascade crossregulation and leads to abnormal hepatogenesis. Proc Natl Acad Sci USA. 1998;95:6881–6886. [PMC free article: PMC22670] [PubMed: 9618507]
- Giroux S, Charron J. Defective development of the embryonic liver in N-myc-deficient mice. Dev Biol. 1998;195:16–28. [PubMed: 9520320]
- Gouon-Evans V, Boussemart L, Gadue P, Nierhoff D, Koehler C.I, Kubo A, Shafritz D.A, Keller G. BMP-4 is required for hepatic specification of mouse embryonic stem cell-derived definitive endoderm. Nat Biotechnol. 2006;24:1402–1411. [PubMed: 17086172]
- Grapin-Botton A. Antero-posterior patterning of the vertebrate digestive tract: 40 years after Nicole Le Douarin's PhD thesis. Int J Dev Biol. 2005;49:335–347. [PubMed: 15906249]
- Gualdi R, Bossard P, Zheng M, Hamada Y, Coleman J.R, Zaret K.S. Hepatic specification of the gut endoderm in vitro: cell signaling and transcriptional control. Genes Dev. 1996;10:1670–1682. [PubMed: 8682297]
- Gunes C, Heuchel R, Georgiev O, Muller K.H, Lichtlen P, Bluthmann H, Marino S, Aguzzi A, Schaffner W. Embryonic lethality and liver degeneration in mice lacking the metal-responsive transcriptional activator MTF-1. Embo J. 1998;17:2846–2854. [PMC free article: PMC1170625] [PubMed: 9582278]
- Hay D.C, Zhao D, Fletcher J, Hewitt Z.A, McLean D, Urruticoechea-Uriguen A, Black J.R, Elcombe C, Ross J.A, Wolf R, Cui W. Efficient Differentiation of Hepatocytes from Human Embryonic Stem Cells Exhibiting Markers Recapitulating Liver Development In Vivo. Stem Cells. 2008 [PubMed: 18238852]
- Hentsch B, Lyons I, Li R, Hartley L, Lints T.J, Adams J. M, Harvey R.P. Hlx homeo box gene is essential for an inductive tissue interaction that drives expansion of embryonic liver and gut. Genes Dev. 1996;10:70–79. [PubMed: 8557196]
- Heo J, Factor V.M, Uren T, Takahama Y, Lee J.S, Major M, Feinstone S.M, Thorgeirsson S.S. Hepatic precursors derived from murine embryonic stem cells contribute to regeneration of injured liver. Hepatology. 2006;44:1478–1486. [PubMed: 17133486]
- Hilberg F, Aguzzi A, Howells N, Wagner E.F. c-jun is essential for normal mouse development and hepatogenesis. Nature. 1993;365:179–181. [PubMed: 8371760]
- Holtzinger A, Evans T. Gata4 regulates the formation of multiple organs. Development. 2005;132:4005–4014. [PubMed: 16079152]
- Horb M.E, Slack J.M. Endoderm specification and differentiation in Xenopus embryos. Dev Biol. 2001;236:330–343. [PubMed: 11476575]
- Horslen S.P, Fox I.J. Hepatocyte transplantation. Transplantation. 2004;77:1481–1486. [PubMed: 15239608]
- Houssaint E. Differentiation of the mouse hepatic primordium. I. An analysis of tissue interactions in hepatocyte differentiation. Cell Differ. 1980;9:269–279. [PubMed: 7438211]
- Hu M, Kurobe M, Jeong Y.J, Fuerer C, Ghole S, Nusse R, Sylvester K.G. Wnt/beta-catenin signaling in murine hepatic transit amplifying progenitor cells. Gastroenterology. 2007;133:1579–1591. [PubMed: 17983805]
- Hunter M.P, Wilson C.M, Jiang X, Cong R, Vasavada H, Kaestner K.H, Bogue C.W. The homeobox gene Hhex is essential for proper hepatoblast differentiation and bile duct morphogenesis. Dev Biol. 2007;308:355–367. [PMC free article: PMC2045067] [PubMed: 17580084]
- Hussain S.Z, Sneddon T, Tan X, Micsenyi A, Michalopoulos G.K, Monga S.P. Wnt impacts growth and differentiation in ex vivo liver development. Exp Cell Res. 2004;292:157–169. [PubMed: 14720515]
- Iida I, Johkura K, Teng R, Kubota S, Cui L, Zhao X, Ogiwara N, Okouchi Y, Asanuma K, Nakayama J, Sasaki K. Immunohistochemical localization of hepatocyte growth factor activator (HGFA) in developing mouse liver tissues. Heterogeneous distribution of HGFA protein. J Histochem Cytochem. 2003;51:1139–1149. [PubMed: 12923239]
- Ijpenberg A, Perez-Pomares J.M, Guadix J.A, Carmona R, Portillo-Sanchez V, Macias D, Hohenstein P, Miles C.M, Hastie N.D, Munoz-Chapuli R. Wt1 and retinoic acid signaling are essential for stellate cell development and liver morphogenesis. Dev Biol. 2007;312:157–170. [PubMed: 18028902]
- Imamura M, Kojima T, Lan M, Son S, Murata M, Osanai M, Chiba H, Hirata K, Sawada N. Oncostatin M induces upregulation of claudin-2 in rodent hepatocytes coinciding with changes in morphology and function of tight junctions. Exp Cell Res. 2007;313:1951–1962. [PubMed: 17434483]
- Ishikawa K.S, Masui T, Ishikawa K, Shiojiri N. Immunolocalization of hepatocyte growth factor and its receptor (c-Met) during mouse liver development. Histochem Cell Biol. 2001;116:453–462. [PubMed: 11735009]
- Ito Y, Matsui T, Kamiya A, Kinoshita T, Miyajima A. Retroviral gene transfer of signaling molecules into murine fetal hepatocytes defines distinct roles for the STAT3 and ras pathways during hepatic development. Hepatology. 2000;32:1370–1376. [PubMed: 11093744]
- Jelnes P, Santoni-Rugiu E, Rasmussen M, Friis S.L, Nielsen J. H, Tygstrup N, Bisgaard H.C. Remarkable heterogeneity displayed by oval cells in rat and mouse models of stem cell-mediated liver regeneration. Hepatology. 2007;45:1462–1470. [PubMed: 17538966]
- Johnson L, Greenbaum D, Cichowski K, Mercer K, Murphy E, Schmitt E, Bronson R.T, Umanoff H, Edelmann W, Kucherlapati R, Jacks T. K-ras is an essential gene in the mouse with partial functional overlap with N-ras. Genes Dev. 1997;11:2468–2481. [PMC free article: PMC316567] [PubMed: 9334313]
- Jung J, Zheng M, Goldfarb M, Zaret K.S. Initiation of mammalian liver development from endoderm by fibroblast growth factors. Science. 1999;284:1998–2003. [PubMed: 10373120]
- Kalinichenko V.V, Zhou Y, Bhattacharyya D, Kim W, Shin B, Bambal K, Costa R.H. Haploinsufficiency of the mouse Forkhead Box f1 gene causes defects in gall bladder development. J Biol Chem. 2002;277:12369–12374. [PubMed: 11809759]
- Kamiya A, Gonzalez F.J. TNF-alpha regulates mouse fetal hepatic maturation induced by oncostatin M and extracellular matrices. Hepatology. 2004;40:527–536. [PubMed: 15349890]
- Kamiya A, Kinoshita T, Ito Y, Matsui T, Morikawa Y, Senba E, Nakashima K, Taga T, Yoshida K, Kishimoto T, Miyajima A. Fetal liver development requires a paracrine action of oncostatin M through the gp130 signal transducer. Embo J. 1999;18:2127–2136. [PMC free article: PMC1171297] [PubMed: 10205167]
- Kamiya A, Kinoshita T, Miyajima A. Oncostatin M and hepatocyte growth factor induce hepatic maturation via distinct signaling pathways. FEBS Lett. 2001;492:90–94. [PubMed: 11248243]
- Kelley-Loughnane N, Sabla G.E, Ley-Ebert C, Aronow B.J, Bezerra J.A. Independent and overlapping transcriptional activation during liver development and regeneration in mice. Hepatology. 2002;35:525–534. [PubMed: 11870364]
- Keng V.W, Yagi H, Ikawa M, Nagano T, Myint Z, Yamada K, Tanaka T, Sato A, Muramatsu I, Okabe M. et al. Homeobox gene Hex is essential for onset of mouse embryonic liver development and differentiation of the monocyte lineage. Biochem Biophys Res Commun. 2000;276:1155–1161. [PubMed: 11027604]
- Kodama Y, Hijikata M, Kageyama R, Shimotohno K, Chiba T. The role of notch signaling in the development of intrahepatic bile ducts. Gastroenterology. 2004;127:1775–1786. [PubMed: 15578515]
- Konopka G, Tekiela J, Iverson M, Wells C, Duncan S.A. Junctional adhesion molecule-A is critical for the formation of pseudocanaliculi and modulates E-cadherin expression in hepatic cells. J Biol Chem. 2007;282:28137–28148. [PubMed: 17623668]
- Krupczak-Hollis K, Wang X, Kalinichenko V.V, Gusarova G. A, Wang I.C, Dennewitz M.B, Yoder H.M, Kiyokawa H, Kaestner K. H, Costa R.H. The mouse Forkhead Box m1 transcription factor is essential for hepatoblast mitosis and development of intrahepatic bile ducts and vessels during liver morphogenesis. Dev Biol. 2004;276:74–88. [PubMed: 15531365]
- Kubo A, Shinozaki K, Shannon J.M, Kouskoff V, Kennedy M, Woo S, Fehling H.J, Keller G. Development of definitive endoderm from embryonic stem cells in culture. Development. 2004;131:1651–1662. [PubMed: 14998924]
- Kumar M, Jordan N, Melton D, Grapin-Botton A. Signals from lateral plate mesoderm instruct endoderm toward a pancreatic fate. Dev Biol. 2003;259:109–122. [PubMed: 12812792]
- Kyrmizi I, Hatzis P, Katrakili N, Tronche F, Gonzalez F.J, Talianidis I. Plasticity and expanding complexity of the hepatic transcription factor network during liver development. Genes Dev. 2006;20:2293–2305. [PMC free article: PMC1553211] [PubMed: 16912278]
- Lavon N, Benvenisty N. Study of hepatocyte differentiation using embryonic stem cells. J Cell Biochem. 2005;96:1193–1202. [PubMed: 16211581]
- Lawson K.A, Pedersen R.A. Cell fate, morphogenetic movement and population kinetics of embryonic endoderm at the time of germ layer formation in the mouse. Development. 1987;101:627–652. [PubMed: 3502998]
- Le Douarin N. Induction de l’endoderme pre-hepatique par le mesoderme de l’aire cardiaque chez l’embryon de poulet. Journal of Embryology & Experimental Morphology. 1964a;12:651–664. [PubMed: 14251477]
- Le Douarin N. Isolement experimental du mesenchyme propre du foie et role morphogene de la composante mesodermique dans l’organogenese hepatique. Journal of Embryology & Experimental Morphology. 1964b;12:141–160. [PubMed: 14155401]
- Le Douarin N.M. An experimental analysis of liver devleopment. Med Biol. 1975;53:427–455. [PubMed: 765644]
- Lee C.S, Friedman J.R, Fulmer J.T, Kaestner K.H. The initiation of liver development is dependent on Foxa transcription factors. Nature. 2005;435:944–947. [PubMed: 15959514]
- Lee C.S, Sund N.J, Behr R, Herrera P.L, Kaestner K.H. Foxa2 is required for the differentiation of pancreatic alpha-cells. Dev Biol. 2005;278:484–495. [PubMed: 15680365]
- Lemaigre F.P. Development of the biliary tract. Mech Dev. 2003;120:81–87. [PubMed: 12490298]
- Li J, Ning G, Duncan S.A. Mammalian hepatocyte differentiation requires the transcription factor HNF-4alpha. Genes Dev. 2000;14:464–474. [PMC free article: PMC316377] [PubMed: 10691738]
- Li L, Krantz I.D, Deng Y, Genin A, Banta A.B, Collins C.C, Qi M, Trask B.J, Kuo W.L, Cochran J. et al. Alagille syndrome is caused by mutations in human Jagged1, which encodes a ligand for Notch1. Nat Genet. 1997;16:243–251. [PubMed: 9207788]
- Li Q, Van Antwerp D, Mercurio F, Lee K.F, Verma I.M. Severe liver degeneration in mice lacking the IkappaB kinase 2 gene. Science. 1999;284:321–325. [PubMed: 10195897]
- Lin X, Jung J, Kang D, Xu B, Zaret K.S, Zoghbi H. Prenylcysteine carboxylmethyltransferase is essential for the earliest stages of liver development in mice. Gastroenterology. 2002;123:345–351. [PubMed: 12105862]
- Loomes K.M, Russo P, Ryan M, Nelson A, Underkoffler L, Glover C, Fu H, Gridley T, Kaestner K.H, Oakey R.J. Bile duct proliferation in liver-specific Jag1 conditional knockout mice: effects of gene dosage. Hepatology. 2007;45:323–330. [PubMed: 17366661]
- Lozier J, McCright B, Gridley T. Notch signaling regulates bile duct morphogenesis in mice. PLoS ONE. 2008;3:e1851. [PMC free article: PMC2266994] [PubMed: 18365007]
- Margagliotti S, Clotman F, Pierreux C.E, Beaudry J.B, Jacquemin P, Rousseau G.G, Lemaigre F.P. The Onecut transcription factors HNF-6/OC-1 and OC-2 regulate early liver expansion by controlling hepatoblast migration. Dev Biol. 2007;311:579–589. [PubMed: 17936262]
- Margagliotti S, Clotman F, Pierreux C.E, Lemoine P, Rousseau G. G, Henriet P, Lemaigre F.P. Role of metalloproteinases at the onset of liver development. Dev Growth Differ. 2008;50:331–338. [PubMed: 18445063]
- Martin M, Gallego-Llamas J, Ribes V, Kedinger M, Niederreither K, Chambon P, Dolle P, Gradwohl G. Dorsal pancreas agenesis in retinoic acid-deficient Raldh2 mutant mice. Dev Biol. 2005;284:399–411. [PubMed: 16026781]
- Martinez Barbera J. P, Clements M, Thomas P, Rodriguez T, Meloy D, Kioussis D, Beddington R.S. The homeobox gene Hex is required in definitive endodermal tissues for normal forebrain, liver and thyroid formation. Development. 2000;127:2433–2445. [PubMed: 10804184]
- Matsui T, Kinoshita T, Hirano T, Yokota T, Miyajima A. STAT3 down-regulates the expression of cyclin D during liver development. J Biol Chem. 2002;277:36167–36173. [PubMed: 12147685]
- Matsui T, Kinoshita T, Morikawa Y, Tohya K, Katsuki M, Ito Y, Kamiya A, Miyajima A. K-Ras mediates cytokine-induced formation of E-cadherin-based adherens junctions during liver development. Embo J. 2002;21:1021–1030. [PMC free article: PMC125879] [PubMed: 11867530]
- Matsumoto K, Miki R, Nakayama M, Tatsumi N, Yokouchi Y. Wnt9a secreted from the walls of hepatic sinusoids is esseantial for morphogenesis, proliferations and glycogen accumulation of chick hepatic epithelium. Dev Biol. 2008 in press . [PubMed: 18513713]
- Matsumoto K, Yoshitomi H, Rossant J, Zaret K.S. Liver organogenesis promoted by endothelial cells prior to vascular function. Science. 2001;294:559–563. [PubMed: 11577199]
- Matthews R.P, Lorent K, Russo P, Pack M. The zebrafish onecut gene hnf-6 functions in an evolutionarily conserved genetic pathway that regulates vertebrate biliary development. Dev Biol. 2004;274:245–259. [PubMed: 15385156]
- McCright B, Lozier J, Gridley T. A mouse model of Alagille syndrome: Notch2 as a genetic modifier of Jag1 haploinsufficiency. Development. 2002;129:1075–1082. [PubMed: 11861489]
- McKenzie T. J, Lillegard J. B, Nyberg S. L. Artificial and bioartificial liver support. Semin Liver Dis. 2008;28:210–217. [PubMed: 18452120]
- McLin V.A, Rankin S.A, Zorn A.M. Repression of Wnt/{beta}-catenin signaling in the anterior endoderm is essential for liver and pancreas development. Development. 2007;134:2207–2217. [PubMed: 17507400]
- Medico E, Gentile A, Lo Celso C, Williams T.A, Gambarotta G, Trusolino L, Comoglio P.M. Osteopontin is an autocrine mediator of hepatocyte growth factor-induced invasive growth. Cancer Res. 2001;61:5861–5868. [PubMed: 11479227]
- Medlock E.S, Haar J.L. The liver hemopoietic environment: I. Developing hepatocytes and their role in fetal hemopoiesis. Anat Rec. 1983;207:31–41. [PubMed: 6638531]
- Michalopoulos G.K. Liver regeneration. J Cell Physiol. 2007;213:286–300. [PMC free article: PMC2701258] [PubMed: 17559071]
- Michalopoulos G.K, Bowen W, Nussler A.K, Becich M.J, Howard T.A. Comparative analysis of mitogenic and morphogenic effects of HGF and EGF on rat and human hepatocytes maintained in collagen gels. J Cell Physiol. 1993;156:443–452. [PubMed: 8360254]
- Michalopoulos G.K, Bowen W.C, Mule K, Luo J. HGF-, EGF-, and dexamethasone-induced gene expression patterns during formation of tissue in hepatic organoid cultures. Gene Expr. 2003;11:55–75. [PMC free article: PMC1913286] [PubMed: 12837037]
- Micsenyi A, Tan X, Sneddon T, Luo J.H, Michalopoulos G.K, Monga S.P. Beta-catenin is temporally regulated during normal liver development. Gastroenterology. 2004;126:1134–1146. [PubMed: 15057752]
- Mikula M, Schreiber M, Husak Z, Kucerova L, Ruth J, Wieser R, Zatloukal K, Beug H, Wagner E.F, Baccarini M. Embryonic lethality and fetal liver apoptosis in mice lacking the c-raf-1 gene. Embo J. 2001;20:1952–1962. [PMC free article: PMC125416] [PubMed: 11296228]
- Monga S.P, Mars W.M, Pediaditakis P, Bell A, Mule K, Bowen W. C, Wang X, Zarnegar R, Michalopoulos G.K. Hepatocyte growth factor induces Wnt-independent nuclear translocation of beta-catenin after Met-beta-catenin dissociation in hepatocytes. Cancer Res. 2002;62:2064–2071. [PubMed: 11929826]
- Monga S.P, Monga H.K, Tan X, Mule K, Pediaditakis P, Michalopoulos G.K. Beta-catenin antisense studies in embryonic liver cultures: role in proliferation, apoptosis, and lineage specification. Gastroenterology. 2003;124:202–216. [PubMed: 12512043]
- Moore-Scott B.A, Opoka R, Lin S.C, Kordich J.J, Wells J.M. Identification of molecular markers that are expressed in discrete anterior-posterior domains of the endoderm from the gastrula stage to mid-gestation. Dev Dyn. 2007;236:1997–2003. [PubMed: 17576135]
- Motoyama J, Kitajima K, Kojima M, Kondo S, Takeuchi T. Organogenesis of the liver, thymus and spleen is affected in jumonji mutant mice. Mech Dev. 1997;66:27–37. [PubMed: 9376320]
- Moumen A, Ieraci A, Patane S, Sole C, Comella J.X, Dono R, Maina F. Met signals hepatocyte survival by preventing Fas-triggered FLIP degradation in a PI3k-Akt-dependent manner. Hepatology. 2007;45:1210–1217. [PubMed: 17464994]
- Nishina H, Vaz C, Billia P, Nghiem M, Sasaki T, De la Pompa J. L, Furlonger K, Paige C, Hui C, Fischer K.D. et al. Defective liver formation and liver cell apoptosis in mice lacking the stress signaling kinase SEK1/MKK4. Development. 1999;126:505–516. [PubMed: 9876179]
- Ober E.A, Verkade H, Field H.A, Stainier D.Y. Mesodermal Wnt2b signalling positively regulates liver specification. Nature. 2006;442:688–691. [PubMed: 16799568]
- Ochsner S.A, Strick-Marchand H, Qiu Q, Venable S, Dean A, Wilde M, Weiss M.C, Darlington G.J. Transcriptional profiling of bipotential embryonic liver cells to identify liver progenitor cell surface markers. Stem Cells. 2007;25:2476–2487. [PMC free article: PMC2853184] [PubMed: 17641245]
- Oda T, Elkahloun A.G, Pike B.L, Okajima K, Krantz I.D, Genin A, Piccoli D.A, Meltzer P.S, Spinner N.B, Collins F.S, Chandrasekharappa S.C. Mutations in the human Jagged1 gene are responsible for Alagille syndrome. Nat Genet. 1997;16:235–242. [PubMed: 9207787]
- Odom D.T, Zizlsperger N, Gordon D.B, Bell G.W, Rinaldi N.J, Murray H.L, Volkert T.L, Schreiber J, Rolfe P.A, Gifford D.K. et al. Control of pancreas and liver gene expression by HNF transcription factors. Science. 2004;303:1378–1381. [PMC free article: PMC3012624] [PubMed: 14988562]
- Oertel M, Menthena A, Chen Y.Q, Teisner B, Jensen C.H, Shafritz D.A. Purification of fetal liver stem/progenitor cells containing all the repopulation potential for normal adult rat liver. Gastroenterology. 2008;134:823–832. [PubMed: 18262526]
- Oertel M, Shafritz D.A. Stem cells, cell transplantation and liver repopulation. Biochim Biophys Acta. 2008;1782:61–74. [PMC free article: PMC2857398] [PubMed: 18187050]
- Okada T.S. Experimental studies on the differentiation of the endodermal organs in Amphibia. II. Differentiating potencies of the presumptive endoderm in the presence of the mesodermal tissues. Mem Coll Univ Kyoto. 1954;21:7–14.
- Okada T.S. Epithelio-mesenchymal relationships in the regional differentiation of the digestive tract in the amphibian embryo. Roux's Arch Dev Biol. 1960;152:1–21. [PubMed: 28354915]
- Papoutsi M, Dudas J, Becker J, Tripodi M, Opitz L, Ramadori G, Wilting J. Gene regulation by homeobox transcription factor Prox1 in murine hepatoblasts. Cell Tissue Res. 2007;330:209–220. [PubMed: 17828556]
- Parviz F, Matullo C, Garrison W.D, Savatski L, Adamson J.W, Ning G, Kaestner K.H, Rossi J.M, Zaret K.S, Duncan S.A. Hepatocyte nuclear factor 4alpha controls the development of a hepatic epithelium and liver morphogenesis. Nat Genet. 2003;34:292–296. [PubMed: 12808453]
- Pelton R.W, Saxena B, Jones M, Moses H.L, Gold L.I. Immunohistochemical localization of TGF beta 1, TGF beta 2, and TGF beta 3 in the mouse embryo: expression patterns suggest multiple roles during embryonic development. J Cell Biol. 1991;115:1091–1105. [PMC free article: PMC2289937] [PubMed: 1955457]
- Pontoglio M, Barra J, Hadchouel M, Doyen A, Kress C, Bach J.P, Babinet C, Yaniv M. Hepatocyte nuclear factor 1 inactivation results in hepatic dysfunction, phenylketonuria, and renal Fanconi syndrome. Cell. 1996;84:575–585. [PubMed: 8598044]
- Porter F.D, Drago J, Xu Y, Cheema S.S, Wassif C, Huang S.P, Lee E, Grinberg A, Massalas J.S, Bodine D. et al. Lhx2, a LIM homeobox gene, is required for eye, forebrain, and definitive erythrocyte development. Development. 1997;124:2935–2944. [PubMed: 9247336]
- Qu X, Lam E, Doughman Y.Q, Chen Y, Chou Y.T, Lam M, Turakhia M, Dunwoodie S.L, Watanabe M, Xu B. et al. Cited2, a coactivator of HNF4alpha, is essential for liver development. Embo J. 2007;26:4445–4456. [PMC free article: PMC2063472] [PubMed: 17932483]
- Rastegar M, Rousseau G.G, Lemaigre F.P. CCAAT/enhancer-binding protein-alpha is a component of the growth hormone-regulated network of liver transcription factors. Endocrinology. 2000;141:1686–1692. [PubMed: 10803577]
- Reimold A.M, Etkin A, Clauss I, Perkins A, Friend D.S, Zhang J, Horton H.F, Scott A, Orkin S.H, Byrne M.C. et al. An essential role in liver development for transcription factor XBP-1. Genes Dev. 2000;14:152–157. [PMC free article: PMC316338] [PubMed: 10652269]
- Roberts D.J, Johnson R.L, Burke A.C, Nelson C.E, Morgan B.A, Tabin C. Sonic hedgehog is an endodermal signal inducing Bmp-4 and Hox genes during induction and regionalization of the chick hindgut. Development. 1995;121:3163–3174. [PubMed: 7588051]
- Rogler L.E. Selective bipotential differentiation of mouse embryonic hepatoblasts in vitro. Am J Pathol. 1997;150:591–602. [PMC free article: PMC1858272] [PubMed: 9033273]
- Rosenfeld M.E, Prichard L, Shiojiri N, Fausto N. Prevention of hepatic apoptosis and embryonic lethality in RelA/TNFR-1 double knockout mice. Am J Pathol. 2000;156:997–1007. [PMC free article: PMC1876833] [PubMed: 10702415]
- Roskams T, Desmet V. Embryology of extra- and intrahepatic bile ducts, the ductal plate. Anat Rec (Hoboken). 2008;291:628–635. [PubMed: 18484608]
- Rossi J.M, Dunn N.R, Hogan B.L, Zaret K.S. Distinct mesodermal signals, including BMPs from the septum transversum mesenchyme, are required in combination for hepatogenesis from the endoderm. Genes Dev. 2001;15:1998–2009. [PMC free article: PMC312750] [PubMed: 11485993]
- Sachs M, Brohmann H, Zechner D, Muller T, Hulsken J, Walther I, Schaeper U, Birchmeier C, Birchmeier W. Essential role of Gab1 for signaling by the c-Met receptor in vivo. J Cell Biol. 2000;150:1375–1384. [PMC free article: PMC2150711] [PubMed: 10995442]
- Sadler K.C, Amsterdam A, Soroka C, Boyer J, Hopkins N. A genetic screen in zebrafish identifies the mutants vps18, nf2 and foie gras as models of liver disease. Development. 2005;132:3561–3572. [PubMed: 16000385]
- Satohisa S, Chiba H, Osanai M, Ohno S, Kojima T, Saito T, Sawada N. Behavior of tight-junction, adherens-junction and cell polarity proteins during HNF-4alpha-induced epithelial polarization. Exp Cell Res. 2005;310:66–78. [PubMed: 16098509]
- Schmelzer E, Wauthier E, Reid L.M. The phenotypes of pluripotent human hepatic progenitors. Stem Cells. 2006;24:1852–1858. [PubMed: 16627685]
- Schmidt C, Bladt F, Goedecke S, Brinkmann V, Zschiesche W, Sharpe M, Gherardi E, Birchmeier C. Scatter factor/hepatocyte growth factor is essential for liver development. Nature. 1995;373:699–702. [PubMed: 7854452]
- Sekhon S.S, Tan X, Micsenyi A, Bowen W.C, Monga S.P. Fibroblast growth factor enriches the embryonic liver cultures for hepatic progenitors. Am J Pathol. 2004;164:2229–2240. [PMC free article: PMC1615755] [PubMed: 15161655]
- Sergi C, Adam S, Kahl P, Otto H.F. Study of the malformation of ductal plate of the liver in Meckel syndrome and review of other syndromes presenting with this anomaly. Pediatr Dev Pathol. 2000;3:568–583. [PubMed: 11000335]
- Sergi C, Kahl P, Otto H.F. Contribution of apoptosis and apoptosis-related proteins to the malformation of the primitive intrahepatic biliary system in Meckel syndrome. Am J Pathol. 2000;156:1589–1598. [PMC free article: PMC1876920] [PubMed: 10793071]
- Serls A.E, Doherty S, Parvatiyar P, Wells J.M, Deutsch G.H. Different thresholds of fibroblast growth factors pattern the ventral foregut into liver and lung. Development. 2005;132:35–47. [PubMed: 15576401]
- Sharma A.D, Cantz T, Vogel A, Schambach A, Haridass D, Iken M, Bleidissel M, Manns M.P, Scholer H.R, Ott M. Murine embryonic stem cell-derived hepatic progenitor cells engraft in recipient livers with limited capacity of liver tissue formation. Cell Transplant. 2008;17:313–323. [PubMed: 18522234]
- Shen M.M. Nodal signaling: developmental roles and regulation. Development. 2007;134:1023–1034. [PubMed: 17287255]
- Shin D, Shin C.H, Tucker J, Ober E.A, Rentzsch F, Poss K.D, Hammerschmidt M, Mullins M.C, Stainier D.Y. Bmp and Fgf signaling are essential for liver specification in zebrafish. Development. 2007;134:2041–2050. [PubMed: 17507405]
- Shiojiri N. The origin of intrahepatic bile duct cells in the mouse. J Embryol Exp Morphol. 1984;79:25–39. [PubMed: 6371179]
- Shiojiri N, Sugiyama Y. Immunolocalization of extracellular matrix components and integrins during mouse liver development. Hepatology. 2004;40:346–355. [PubMed: 15368439]
- Shiojiri N, Takeshita K, Yamasaki H, Iwata T. Suppression of C/EBP alpha expression in biliary cell differentiation from hepatoblasts during mouse liver development. J Hepatol. 2004;41:790–798. [PubMed: 15519652]
- Soriano H.E, Bilyeu T.A, Juan T.S, Zhao W, Darlington G.J. DNA binding by C/EBP proteins correlates with hepatocyte proliferation. In Vitro Cell Dev Biol Anim. 1995;31:703–709. [PubMed: 8564082]
- Sosa-Pineda B, Wigle J.T, Oliver G. Hepatocyte migration during liver development requires Prox1. Nat Genet. 2000;25:254–255. [PubMed: 10888866]
- Soto-Gutierrez A, Kobayashi N, Rivas-Carrillo J.D, Navarro-Alvarez N, Zhao D, Okitsu T, Noguchi H, Basma H, Tabata Y, Chen Y. et al. Reversal of mouse hepatic failure using an implanted liver-assist device containing ES cell-derived hepatocytes. Nat Biotechnol. 2006;24:1412–1419. [PubMed: 17086173]
- Soto-Gutierrez A, Navarro-Alvarez N, Zhao D, Rivas-Carrillo J.D, Lebkowski J, Tanaka N, Fox I.J, Kobayashi N. Differentiation of mouse embryonic stem cells to hepatocyte-like cells by co-culture with human liver nonparenchymal cell lines. Nat Protoc. 2007;2:347–356. [PubMed: 17406596]
- Stafford D, Hornbruch A, Mueller P.R, Prince V.E. A conserved role for retinoid signaling in vertebrate pancreas development. Dev Genes Evol. 2004;214:432–441. [PubMed: 15322880]
- Stenvers K.L, Tursky M.L, Harder K.W, Kountouri N, Amatayakul-Chantler S, Grail D, Small C, Weinberg R.A, Sizeland A.M, Zhu H.J. Heart and liver defects and reduced transforming growth factor beta2 sensitivity in transforming growth factor beta type III receptor-deficient embryos. Mol Cell Biol. 2003;23:4371–4385. [PMC free article: PMC156130] [PubMed: 12773577]
- Strick-Marchand H, Morosan S, Charneau P, Kremsdorf D, Weiss M.C. Bipotential mouse embryonic liver stem cell lines contribute to liver regeneration and differentiate as bile ducts and hepatocytes. Proc Natl Acad Sci USA. 2004;101:8360–8365. [PMC free article: PMC420399] [PubMed: 15155906]
- Strick-Marchand H, Weiss M.C. Inducible differentiation and morphogenesis of bipotential liver cell lines from wild-type mouse embryos. Hepatology. 2002;36:794–804. [PubMed: 12297826]
- Strick-Marchand H, Weiss M.C. Embryonic liver cells and permanent lines as models for hepatocyte and bile duct cell differentiation. Mech Dev. 2003;120:89–98. [PubMed: 12490299]
- Suchy F.J. Clinical problems with developmental anomalies of the biliary tract. Semin Gastrointest Dis. 2003;14:156–164. [PubMed: 14719766]
- Suksaweang S, Lin C.M, Jiang T.X, Hughes M.W, Widelitz R.B, Chuong C.M. Morphogenesis of chicken liver: identification of localized growth zones and the role of beta-catenin/Wnt in size regulation. Dev Biol. 2004;266:109–122. [PMC free article: PMC4376314] [PubMed: 14729482]
- Suzuki A, Iwama A, Miyashita H, Nakauchi H, Taniguchi H. Role for growth factors and extracellular matrix in controlling differentiation of prospectively isolated hepatic stem cells. Development. 2003;130:2513–2524. [PubMed: 12702664]
- Suzuki A, Sekiya S, Buscher D, Izpisua Belmonte J.C, Taniguchi H. Tbx3 controls the fate of hepatic progenitor cells in liver development by suppressing p19ARF expression. Development. 2008;135:1589–1595. [PubMed: 18356246]
- Suzuki T, Kanai Y, Hara T, Sasaki J, Sasaki T, Kohara M, Maehama T, Taya C, Shitara H, Yonekawa H. et al. Crucial role of the small GTPase ARF6 in hepatic cord formation during liver development. Mol Cell Biol. 2006;26:6149–6156. [PMC free article: PMC1592812] [PubMed: 16880525]
- Takata N. The differentiation in vivo of the isolated endoderm under the influence of the mesoderm in Triturus Pyrrhogaster. Embryologica. 1960;5:38–70.
- Tan X, Apte U, Micsenyi A, Kotsagrelos E, Luo J.H, Ranganathan S, Monga D.K, Bell A, Michalopoulos G.K, Monga S.P. Epidermal growth factor receptor: a novel target of the Wnt/beta-catenin pathway in liver. Gastroenterology. 2005;129:285–302. [PMC free article: PMC1821080] [PubMed: 16012954]
- Tan X, Behari J, Cieply B, Michalopoulos G.K, Monga S.P. Conditional Deletion of beta-Catenin Reveals Its Role in Liver Growth and Regeneration. Gastroenterology. 2006;131:1561–1572. [PubMed: 17101329]
- Tan X, Yuan Y, Zeng G, Apte U, Thompson M.D, Cieply B, Stolz D.B, Michalopoulos G.K, Kaestner K.H, Monga S.P. beta-Catenin deletion in hepatoblasts disrupts hepatic morphogenesis and survival during mouse development. Hepatology. 2008;47:1667–1679. [PMC free article: PMC4449338] [PubMed: 18393386]
- Tang Y, Katuri V, Dillner A, Mishra B, Deng C.X, Mishra L. Disruption of transforming growth factor-beta signaling in ELF beta-spectrin-deficient mice. Science. 2003;299:574–577. [PubMed: 12543979]
- Tanimizu N, Miyajima A. Notch signaling controls hepatoblast differentiation by altering the expression of liver-enriched transcription factors. J Cell Sci. 2004;117:3165–3174. [PubMed: 15226394]
- Tatsumi N, Miki R, Katsu K, Yokouchi Y. Neurturin-GFRalpha2 signaling controls liver bud migration along the ductus venosus in the chick embryo. Dev Biol. 2007;307:14–28. [PubMed: 17509555]
- Teratani T, Yamamoto H, Aoyagi K, Sasaki H, Asari A, Quinn G, Terada M, Ochiya T. Direct hepatic fate specification from mouse embryonic stem cells. Hepatology. 2005;41:836–846. [PubMed: 15742390]
- Thomas P.Q, Brown A, Beddington R.S. Hex: a homeobox gene revealing peri-implantation asymmetry in the mouse embryo and an early transient marker of endothelial cell precursors. Development. 1998;125:85–94. [PubMed: 9389666]
- Tiso N, Filippi A, Pauls S, Bortolussi M, Argenton F. BMP signalling regulates anteroposterior endoderm patterning in zebrafish. Mech Dev. 2002;118:29–37. [PubMed: 12351167]
- Tomizawa M, Garfield S, Factor V, Xanthopoulos K. G. Hepatocytes deficient in CCAAT/enhancer binding protein alpha (C/EBP alpha) exhibit both hepatocyte and biliary epithelial cell character. Biochem Biophys Res Commun. 1998;249:1–5. [PubMed: 9705820]
- Tremblay K.D, Zaret K.S. Distinct populations of endoderm cells converge to generate the embryonic liver bud and ventral foregut tissues. Dev Biol. 2005;280:87–99. [PubMed: 15766750]
- Wandzioch E, Kolterud A, Jacobsson M, Friedman S.L, Carlsson L. Lhx2-/- mice develop liver fibrosis. Proc Natl Acad Sci USA. 2004;101:16549–16554. [PMC free article: PMC526277] [PubMed: 15536133]
- Wang N.D, Finegold M.J, Bradley A, Ou C.N, Abdelsayed S.V, Wilde M.D, Taylor L.R, Wilson D.R, Darlington G.J. Impaired energy homeostasis in C/EBP alpha knockout mice. Science. 1995;269:1108–1112. [PubMed: 7652557]
- Wang Z, Dolle P, Cardoso W.V, Niederreither K. Retinoic acid regulates morphogenesis and patterning of posterior foregut derivatives. Dev Biol. 2006;297:433–445. [PubMed: 16806149]
- Watanabe T, Nakagawa K, Ohata S, Kitagawa D, Nishitai G, Seo J, Tanemura S, Shimizu N, Kishimoto H, Wada T. et al. SEK1/MKK4-mediated SAPK/JNK signaling participates in embryonic hepatoblast proliferation via a pathway different from NF-kappaB-induced anti-apoptosis. Dev Biol. 2002;250:332–347. [PubMed: 12376107]
- Watt A.J, Garrison W.D, Duncan S.A. HNF4: a central regulator of hepatocyte differentiation and function. Hepatology. 2003;37:1249–1253. [PubMed: 12774000]
- Watt A.J, Zhao R, Li J, Duncan S.A. Development of the mammalian liver and ventral pancreas is dependent on GATA4. BMC Dev Biol. 2007;7:37. [PMC free article: PMC1877807] [PubMed: 17451603]
- Weinstein M, Monga S.P, Liu Y, Brodie S.G, Tang Y, Li C, Mishra L, Deng C.X. Smad proteins and hepatocyte growth factor control parallel regulatory pathways that converge on beta1-integrin to promote normal liver development. Mol Cell Biol. 2001;21:5122–5131. [PMC free article: PMC87237] [PubMed: 11438667]
- Wells J.M, Melton D.A. Early mouse endoderm is patterned by soluble factors from adjacent germ layers. Development. 2000;127:1563–1572. [PubMed: 10725233]
- Yamasaki H, Sada A, Iwata T, Niwa T, Tomizawa M, Xanthopoulos K.G, Koike T, Shiojiri N. Suppression of C/EBPalpha expression in periportal hepatoblasts may stimulate biliary cell differentiation through increased Hnf6 and Hnf1b expression. Development. 2006;133:4233–4243. [PubMed: 17021047]
- Yanai M, Tatsumi N, Hasunuma N, Katsu K, Endo F, Yokouchi Y. FGF signaling segregates biliary cell-lineage from chick hepatoblasts cooperatively with BMP4 and ECM components in vitro. Dev Dyn. 2008;237:1268–1283. [PubMed: 18393311]
- Yasunaga M, Tada S, Torikai-Nishikawa S, Nakano Y, Okada M, Jakt L.M, Nishikawa S, Chiba T, Era T. Induction and monitoring of definitive and visceral endoderm differentiation of mouse ES cells. Nat Biotechnol. 2005;23:1542–1550. [PubMed: 16311587]
- Yovchev M.I, Grozdanov P.N, Zhou H, Racherla H, Guha C, Dabeva M.D. Identification of adult hepatic progenitor cells capable of repopulating injured rat liver. Hepatology. 2008;47:636–647. [PubMed: 18023068]
- Yuan Z.R, Kobayashi N, Kohsaka T. Human Jagged 1 mutants cause liver defect in Alagille syndrome by overexpression of hepatocyte growth factor. J Mol Biol. 2006;356:559–568. [PubMed: 16403414]
- Zaret K.S. Genetic programming of liver and pancreas progenitors: lessons for stem-cell differentiation. Nat Rev Genet. 2008 [PubMed: 18398419]
- Zeng G, Awan F, Otruba W, Muller P, Apte U, Tan X, Gandhi C, Demetris A.J, Monga S.P. Wnt’er in liver: expression of Wnt and frizzled genes in mouse. Hepatology. 2007;45:195–204. [PubMed: 17187422]
- Zhang W, Yatskievych T.A, Baker R.K, Antin P.B. Regulation of Hex gene expression and initial stages of avian hepatogenesis by Bmp and Fgf signaling. Dev Biol. 2004;268:312–326. [PubMed: 15063170]
- Zhao R, Duncan S.A. Embryonic development of the liver. Hepatology. 2005;41:956–967. [PubMed: 15841465]
- Zhao R, Watt A.J, Battle M.A, Li J, Bondow B.J, Duncan S.A. Loss of both GATA4 and GATA6 blocks cardiac myocyte differentiation and results in acardia in mice. Dev Biol. 2008;317:614–619. [PMC free article: PMC2423416] [PubMed: 18400219]
- Zhao R, Watt A.J, Li J, Luebke-Wheeler J, Morrisey E.E, Duncan S.A. GATA6 is essential for embryonic development of the liver but dispensable for early heart formation. Mol Cell Biol. 2005;25:2622–2631. [PMC free article: PMC1061656] [PubMed: 15767668]
- Zorn A.M, Wells J.M. Molecular basis of vertebrate endoderm development. Int Rev Cytol. 2007;259:49–111. [PubMed: 17425939]
- *
Edited by Alexander F. Schier. Last revised October 2, 2008. PublishedOctober 31, 2008. This chapter should be cited as: Zorn, A.M., Liver development (October 31, 2008), StemBook, ed. The Stem Cell Research Community, StemBook, doi/10.3824/stembook.1.25.1, http://www.stembook.org.
Publication Details
Author Information and Affiliations
Authors
Aaron M Zorn1, .
.Affiliations
 To whom correspondence should be addressed. E-mail: Aaron.zorn@chmcc.org
To whom correspondence should be addressed. E-mail: Aaron.zorn@chmcc.orgPublication History
Published October 31, 2008.
Copyright
This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
Publisher
Harvard Stem Cell Institute, Cambridge (MA)
NLM Citation
Zorn AM. Liver development. 2008 Oct 31. In: StemBook [Internet]. Cambridge (MA): Harvard Stem Cell Institute; 2008-. doi: 10.3824/stembook.1.25.1