The content of this book is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 4.0 Unported license. To view the terms and conditions of this license, visit https://creativecommons.org/licenses/by-nc-nd/4.0/
NCBI Bookshelf. A service of the National Library of Medicine, National Institutes of Health.
Varki A, Cummings RD, Esko JD, et al., editors. Essentials of Glycobiology [Internet]. 4th edition. Cold Spring Harbor (NY): Cold Spring Harbor Laboratory Press; 2022. doi: 10.1101/glycobiology.4e.3
This chapter discusses the diversity in structure and properties that results when multiple monosaccharides (Chapter 2) are linked together to form oligosaccharides and polysaccharides (the latter comprising much of the biomass on the planet). Some examples of the more complex polymeric assemblies that occur in nature are presented, and how these remarkable structures are generated is discussed.
GLYCANS IN NATURE ARE OFTEN CONJUGATED
Except in their roles as sources of energy for living organisms, sugars seldom occur in nature as monosaccharides. Instead, they serve as building blocks for more complex molecules. In the most common process, an initial sugar is linked to an aglycone (often a lipid or a protein) and this sugar is further elaborated by covalently joining other sugars through glycosidic linkages (Chapter 2) between the anomeric carbon of the sugar being added and a hydroxyl oxygen of an existing sugar. The resulting glycans are called oligosaccharides (usually less than a dozen monosaccharides) or polysaccharides (usually more than a dozen monosaccharides). The latter are usually built on a core of repeating subunits of linked monosaccharides. The way in which assembly of both oligosaccharides and polysaccharides occurs produces structures of enormous diversity and widely varying properties. This allows glycans to fill roles that vary from cell-surface interactions with proteins important in differentiation, recognition, and proliferation of cells, to interactions with other glycans that generate the mechanical properties of plant and microbial cell walls.
DIVERSITY FROM OLIGOSACCHARIDE BRANCHING
Diverse structures can be created by simply linking different monosaccharides through glycosidic bonds, to make oligosaccharides or polysaccharides. The diversity arises not only from the choice of sugars but also from the way they are linked. If there were just one way to link monosaccharides, the choice among the dozen or so commonly used sugars would make the resulting polysaccharides more diverse than polynucleotides (four nucleotide choices for DNA and RNA) but less diverse than polypeptides (20 amino acid choices for mammalian proteins). However, the possibility of making glycosidic bonds between the anomeric carbon of one sugar and any one of the unmodified hydroxyl groups in another mono- or oligosaccharide adds to the diversity, by allowing not only more linear products, but also branched products in which more than one hydroxyl group on a given sugar is used to make glycosidic bonds. In addition, each anomeric carbon is a stereogenic center, and therefore each glycosidic linkage can be constructed having either the α- or β-configuration. Building an oligosaccharide, such as a tetrasaccharide (four sugars) with an unlinked reducing end, using only a single sugar in one ring form, such as glucopyranose, the authors could construct 1792 distinct structures. Of course, not all theoretically possible molecules are produced in nature because of the lack of enzymes to build them, but many are made contributing to the wide range of functional properties that allow carbohydrates to play many important roles.
Branching is a prime characteristic of many glycans found on mammalian cell surfaces. Glycans representing two major types of eukaryotic protein glycosylation are shown in Figure 3.1. An N-glycan makes a glycosidic bond with the side-chain nitrogen of an asparagine residue that is a part of a consensus peptide sequence NX(S/T). An O-glycan makes a glycosidic bond with the terminal oxygen of a serine or threonine residue. N-Glycans contain a core composed of three mannose residues and two N-acetylglucosamine residues (Manα1-6[Manα1-3]Manβ1-4GlcNAcβ1-4GlcNAcβ1-N-Asn). The depicted glycan is a biantennary glycan with branches linked at the 3- and 6-positions of the first mannose residue in the glycan chain. However, more complex structures exist with three and four branches. Details of the synthesis and biological importance of these glycans are presented in Chapter 9. The depicted O-glycan contains a typical core structure (one of four common cores) that begins at the reducing end with an N-acetylgalactosamine α-linked to a serine or threonine (GlcNAcβ1-6[Galβ1-3]GalNAcα1-O-Ser/Thr). It begins as a biantennary structure but can be further extended toward the nonreducing end to form more complex structures. The synthesis and biological importance of O-glycans are described in Chapters 10 (O-GalNAc) and 19 (O-GlcNAc).

FIGURE 3.1.
Examples of branched structures in N- and O-linked glycans.
Both glycans depicted are terminated with a sialic acid (often Neu5Ac in humans) at their nonreducing end. Sialylation is characteristic of mammalian glycans and important for immune response. In protein–glycan interactions, not only are particular residues recognized, but often their position in a branching structure is recognized as well. An interesting example is the interaction of the enzyme that adds the 2-6-linked sialic acids to the termini of N-glycans. It has a high preference for adding to the 1-3-linked branch (by more than an order of magnitude) despite the fact that the residues on both branches are identical back to the mannose at the branch point (Galβ1-4GlcNAcβ1-2Manα-). This may give some indication of the extent to which branching plays a role in recognition processes.
Unique among mammals are free oligosaccharides present in milk. Human milk oligosaccharides (HMOs) are highly abundant in breast milk. Unlike most glycans that are conjugated to either proteins or lipids, HMOs are unconjugated in their native states. Thus, they contain a reactive aldehyde on the reducing end. HMOs decrease during lactation but are often even more abundant than proteins. Although the precise details of their synthesis are still not well known, HMO structures are more similar to glycans on glycolipids and O-glycans on proteins. The structures are based on a lactose core (Galβ1-4Glc) and extend either in branched or linear forms. Extensions are composed primarily of glucose, galactose, N-acetylglucosamine, fucose, and sialic acids. As molecules, they are generally small, with the majority of the structures in the range of three to six monosaccharides; however, structures with more than 20 monosaccharides have been observed. HMO structures have been characterized by advanced glycomic analytical tools such as liquid chromatography–mass spectrometry (LC-MS) (see Chapter 50). Hundreds of structures have been observed, although one mother typically produces about 100 structures.
There are two HMO phenotypes (from four genotypes) that determine the types of structures in the milk. Mothers who secrete their blood type or the Lewis b epitope are termed secretors. Secretors produce structures that contain α1-2-linked fucose. Nonsecretors produce little or no α1-2-linked fucose but instead produce fucosylated glycans that are primarily (1-3)- or (1-4)-linked. One of the most abundant components in milk of secretor mothers is 2-fucosyllactose, whereas nonsecretor mothers produce very little or none of this compound. Similarly, secretor milk is slightly richer in fucosylated compounds, whereas nonsecretor milk is slightly richer in sialylated compounds.
HMOs have altered our concept of the function of food. Human milk was thought to be primarily nutritive. However, despite the large abundances of HMOs in the milk, there are no human enzymes in the infant gut that can degrade them. Instead, characterization of the gut microbiome of infants have revealed that HMOs are primarily food for the gut bacteria. Indeed, Bifidobacteria, found in the microbiome of heathy breastfed infants, have glycosyl hydrolases that specifically catabolize HMOs. Additionally, HMOs may also have a protective role by blocking the binding of pathogens in the gut.
STRUCTURAL AND STORAGE POLYSACCHARIDES
Linkage variation plays an important role in the structural properties of polysaccharides as illustrated for two closely related glucose polymers having repeating units (RUs) of -[4Glcβ1-]n and -[4Glcα1-]n. The former is the structural polymer, cellulose, that forms the foundation of all plant cell walls and is a major component of materials such as wood and cotton. The latter is starch, an easily digestible material with no significant structural utility. The ability of one polymer to associate in long fibrils having both crystalline and amorphous regions contributing to its structural properties, and the lack of these tendencies in the other polymer, is clearly linked to the stereochemistry at the anomeric carbon (β for cellulose and α for starch) and the preferred torsional angles about the C1′-O4 and O4-C4 bonds of the glycosidic linkages. The torsion angles are referred to as φ and ψ, much the same as primary structural variables in polypeptides; by International Union of Pure and Applied Chemistry (IUPAC) convention they are defined by four connected atoms, O5′-C1′-O4-C4 and C1′-O4-C4-C3, respectively. Because the torsion angle can be monitored directly through couplings between protons at the ends of the glycosidic linkage observed in nuclear magnetic resonance (NMR) spectra, an alternate NMR definition is also in common use—namely, H1′-C1′-O4-C4 for φ and C1′-O4-C4-H4 for ψ. Glycosidic torsion angles differ considerably in crystalline cellulose and starch. Using IUPAC definitions they prefer ∼−95° and ∼+95° in the former and ∼+115° and ∼+120° in the latter. Figure 3.2 depicts these differences in the repeating disaccharide units of cellulose and starch as represented by the isolated disaccharides having common names cellobiose and maltose, respectively. These local preferences influence association properties and ultimately structural characteristics. When extended to a long cellulose polymer, cellobiose units generate elongated strands that can pack and interact with other strands through hydrogen bonds to form layers. Layers in turn interact through a combination of forces to form fibrils composed of 18 polymer chains. The more helical strands in starch cannot pack easily and result in a more amorphous material.
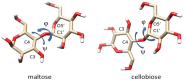
FIGURE 3.2.
Repeating units from cellulose and starch showing conformation and glycosidic torsion angles φ and ψ.
Other important polysaccharides, such as pectins found in plant cell walls, help plants accommodate changes during growth. Pectins are some of the most complex polymers in nature. They are based on polymers of α(1-4)-linked galacturonic acid (GalA) or the RU, -2)-α-L-Rha-(1-4)-α-D-GalA-(1-, and may contain additional sugar and nonsugar substituents, including methyl and acetyl esters. The negatively charged carboxyl group at the 6-position of the GalA residues contributes to the water solubility of these polymers and the long-range interactions that give them gelling properties useful in the food industry. Oligo- and polysaccharides of plant origin are discussed in detail in Chapter 24.
Animals also use polysaccharides for various purposes. Glycogen is a storage polymer related to starch in that it is a glucose polymer with primarily α(1-4)-linkages connecting glucose residues, but it is highly branched having additional α(1-6)-linkages to some of the glucose residues. Structural polymers also exist; for example, the repeating polymer of N-acetylglucosamine, -[4GlcNAcβ1-]n, is the primary component of chitin, the material that forms the exoskeletons of arachnids, crustaceans, and insects (Chapter 26). Modifications of glucose residues by replacing the hydroxyl group at 2-position with an amine group and subsequent N-acetylation change the structural properties significantly. These changes allow for the formation of composites with proteins and minerals that lead to additional variation in structure and function.
CELL-SURFACE POLYSACCHARIDES OF ANIMALS
Most cell-surface polysaccharides found in animals belong to a class of glycans known as glycosaminoglycans (GAGs) (Chapter 17). Abundantly present on the cell surface as well as in the extracellular matrix, GAGs are linear macromolecules with molecular mass of >15,000 Da. The building blocks of most GAGs are composed of an amino-substituted sugar and a hexuronic acid residue. Modifications on the sugar residues—in particular, the sulfation of hydroxyl or the amino groups—are common. The sulfates and hexuronic acid carboxylate groups are negatively charged under physiological conditions. Thus, GAGs are the most anionic molecules present in the animal kingdom. Commonly found GAGs include chondroitin sulfate, dermatan sulfate, heparan sulfate (HS), hyaluronic acid, and keratan sulfate. These GAGs structurally differ in their disaccharide RUs. For example, chondroitin sulfates consist of disaccharide RUs of [4GlcAβ1-3GalNAcβ1-]n, whereas HS consists of disaccharide RUs of [4GlcAβ1-4GlcNAcα1-]n or [4IdoAα1-4GlcNSα1-]n (Figure 3.3A). Structural diversity of these polymers is primarily a result of additional sulfation of hydroxyl groups and is discussed below for the diverse HS polymers found in mammals. GAGs found in other animals can be distinct from mammalian GAGs by virtue of further modifications. For example, marine invertebrates can carry particularly unique sulfation patterns (i.e., 3-O-sulfation on GlcA residues) and distinct side-chain modifications (i.e., fucosylation on chondroitin sulfate).
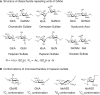
FIGURE 3.3.
Structures of (A) disaccharide repeating units of different glycosaminoglycans and (B) the conformations of monosaccharides from heparan sulfate.
Structure–Function Relationships of Animal Polysaccharides
GAGs show biological function primarily through their interactions with the hundreds of GAG-binding proteins found on cellular surfaces and in extracellular spaces. The structural factors that affect the strength and specificity of binding are key to the elicitation of a proper biological response. HS, the most studied member of the GAG family, provides a good illustration of the wide range of physiological and pathophysiological functions affected. HS, for example, is involved in trimolecular interactions with growth factors and growth factor receptors, and thereby participates in regulating embryonic development. HS interacts with proteases and protease inhibitors in the blood to control the coagulation process and binds to viral envelope proteins as a receptor for viral infections. Also, heparin, a highly sulfated form of HS, is a commonly used anticoagulant drug in the clinic. A more complete discussion of the biological function of HS can be found in Chapters 17 and Chapter 38. This section uses the interaction between HS and proteins to illustrate how structural factors affect binding between GAGs and proteins in general.
One factor that contributes to the structural diversity of HS-protein interactions is the conformational flexibility of L-IdoA and its derivative, L-IdoA2S (2-O-sulfo iduronic acid). Present in the pyranose form, the six-membered ring structure of IdoA or IdoA2S can adopt either chair or skew-boat conformations (Figure 3.3B). Until now, only the 4C1 chair conformation has been verified experimentally for GlcA and GlcN in HS, but both the 1C4 chair and the 2S0 conformations have been found for IdoA2S residues in crystal structures containing HS. In solution, NMR studies show IdoA2S and IdoA residues to be present in a mixture of 1C4 and 2S0 conformations. The conformational flexibility in IdoA residues likely allows orientation of the sulfate groups in HS to maximize the binding affinity to proteins. Structural elements that dictate preferences for one form over the other, including the possible effects from the sulfated monosaccharide sequences around the IdoA/IdoA2S residue, are subject to further investigation.
A second factor contributing to structural diversity and selective interactions with certain HS-binding proteins is the size of the sulfated saccharide domains. HS isolated from natural sources shows domain-like structures, with clusters of six to eight saccharides forming the highly sulfated domains known as S-domains. These regions are separated by nonsulfated saccharide residues, consisting of GlcA-GlcNAc RUs, known as NAc-domains. The S-domains contain primarily IdoA residues that likely contribute the flexibility needed to optimize binding to proteins and elicit desired biological activities. The contribution of NAc-domains to the functions of HS has not been fully established. However, a possible role may be to appropriately position S-domains in a single polysaccharide chain for interaction with multiple proteins. One example is the interaction of heparin with antithrombin and thrombin. In this complex, one part of the heparin chain interacts with antithrombin, and another part of the heparin chain interacts with thrombin. Between the antithrombin-binding domain and thrombin-binding domain, a linker of six to seven saccharide residues exists.
Cellular Regulation of the Structures of GAGs
Unlike proteins and nucleic acids, the biosynthesis of GAGs is not regulated by a well-defined template. Instead, each member of the GAG family is synthesized by a unique pathway (Chapter 17). The HS synthesis pathway, for example, involves multiple enzymes, including specific glycosyltransferases (or HS polymerase), an epimerase, and several sulfotransferases. Moreover, HS is biosynthesized as a polymer covalently linked to a proteoglycan that consists of a core protein and polysaccharide side chains; the functions of these proteoglycans are, in fact, dominated by the properties of the HS chains added. Although this is a non-template-driven process, the overall structures of HS generally remain unchanged between generations. There is considerable interest in understanding the mechanism that controls the structure of HS.
The biosynthesis of chondroitin sulfate and dermatan sulfate is less understood than that of HS. Chondroitin sulfate tends to be synthesized on different proteoglycan core proteins and requires a different set of polymerases and sulfotransferases than the enzymes specifically recognizing HS polysaccharides. To synthesize the IdoA residues in dermatan sulfate, a specialized epimerase is required. Hyaluronic acid biosynthesis is quite different: it is not synthesized on a core protein, it does not take place in the endoplasmic reticulum and Golgi, and it only requires one hyaluronic acid synthase (a dual activity glycosyltransferase), as the polysaccharide does not contain sulfate groups or IdoA residues (Chapter 16).
BACTERIAL POLYSACCHARIDES
The interactions of bacteria with their environment provide an excellent example of how polysaccharide properties play an important role in an organism's survival. Bacterial polysaccharides are especially diverse, in that they can include a larger number of distinct sugar residues in their RUs (usually two to six residues), and they can include branching. Many of them are parts of bacterial cell membranes where they serve important structural and protective roles. Because of their location on the outside of the cell, bacterial polysaccharides such as lipopolysaccharides (LPSs), capsular polysaccharides (CPSs), and exopolysaccharides (EPSs) are often potent antigens that elicit a strong immune response in humans. LPSs carry long polysaccharides called O-antigens and are unique to Gram-negative bacteria where they constitute the major component of the outer leaflet of the outer membrane. These types of bacteria may also carry CPSs forming a relatively dense additional layer around the bacterial cell. Cell wall lipoteichoic acids and teichoic acids are unique components of Gram-positive bacteria, which are often surrounded by a CPS or a less dense EPS layer.
The variation in biosynthetic pathways among bacterial species is ultimately responsible for the diversity in bacterial polysaccharides. The biosynthesis of bacterial polysaccharides is discussed further in Chapter 21. Here the authors present a few examples to illustrate how synthesis of backbone structures, branching, and postpolymerization modification lead to a diverse set of polymer structures. In one of the biosynthetic pathways, sugar residues are added sequentially onto an anchor molecule; thus, the polymer is growing from the terminal, nonreducing end until a termination entity/substituent is added that precludes further chain elongation—for example, in the O-antigen polysaccharide of the LPS from Escherichia coli O8 (Figure 3.4A). The polymer is linear, and although sugar residues are added step-by-step to form the polysaccharide, RUs can be identified.
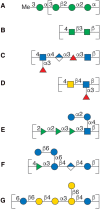
FIGURE 3.4.
Schematic representation in SNFG (Symbol Nomenclature for Glycans) format of repeating units of bacterial polysaccharides: (A) O-antigen of E. coli O8, (B) O-polysaccharide of Salmonella enterica O54, (C) O-antigen of E. coli O168, (D) O-polysaccharide (more...)
Some heteropolysaccharides contain two alternating sequentially added sugar residues and a formal RU can be identified. Often a processive glycosyltransferase is responsible for forming this pattern as is the case in the synthesis of the O-polysaccharide of Salmonella enterica O54 (Figure 3.4B). When two sugar residues alternate, branched structures can be formed, as one sugar may give rise to the polymer backbone and one a side chain.
Polysaccharides are not always built from the nonreducing end, and preformed subunits can be used in the assembly. For example, synthesis of the E. coli antigens O5ab and O5ac relies on a preformed linear oligosaccharide, with five sugar residues constituting the RU. The oligosaccharide is built on an undecaprenyl pyrophosphoryl glycoside anchor molecule. This oligosaccharide is then added onto another oligosaccharide–lipid anchor to grow the polymer from the “reducing end.” When polymerization is taking place at the penultimate sugar of the oligosaccharide a branched RU results (cf. O-antigen of E. coli O168 [Figure 3.4C]).
Branching can be introduced by the addition of sugars following polymer backbone formation. Helicobacter pylori O-antigens often contain human blood group structures as part of their RUs. N-acetyl-D-glucosamine and D-galactose are added in a processive manner onto an undecaprenyl pyrophosphoryl carrier, thereby forming a linear polysaccharide composed of Galβ1-4GlcNAc (LacNAc) disaccharide RUs. Subsequently, L-fucosyl residues are added onto the backbone polysaccharide resulting in branched Lewis-type structures (Figure 3.4D).
Bacterial saccharides consist of diverse monosaccharides (Chapter 2) including 6-deoxy-hexoses like L-rhamnose or L-fucose as well as rare sugars that are often found at the terminal position of the oligosaccharide–lipid acceptors. On polymerization, the rare monosaccharides become part of the side chain in branched structures and give rise to structural epitopes that are characteristic for bacterial species. In some cases, these epitopes are the basis for molecular mimicry (Chapter 42). Furthermore, these sugar residues constitute the terminal entity of the polysaccharide chain, which can be recognized by antibodies of the immune system. The polysaccharide may also be decorated by substituents such as amino acids, O-acyl, or phosphodiester groups. It is not uncommon to find O-acetyl groups at the branch-point sugar residue, thereby leading to a highly crowded substitution pattern in which all positions on the sugar residue are either part of a glycosidic linkage or carry a nonsugar substituent. Examples of diversity by substituent addition are the O-antigens of Shigella flexneri in which glucosyl, O-acetyl, or phosphoethanolamine groups are added to the backbone (Figure 3.4E).
Branching and substituent addition affect the properties of polysaccharide solutions, such as gelling and high viscosity. Some polymers are known under their commercial names gellan, welan, and rhamsan (S-194) (Figure 3.4F). The differences among these and similar types of polymers are based on acyl substituents such as O-acetyl or O-succinyl groups and side chains consisting of l-Rha/l-Man, di-glucosyl or di-rhamnosyl groups, as well as an l-Rha/l-Man backbone modification.
The size of polysaccharides produced by bacteria can vary widely. Whereas the O-antigens in LPSs have less than 100 RUs, CPSs and EPSs have higher numbers of RUs (103–105). Branched structures are often present in these polysaccharides (Figure 3.4G) with either more than one sugar residue in the side chain or with two branches within the RU. More information on these complex materials is presented in Chapter 42.
Charge can significantly affect polysaccharide properties. Charged polysaccharides form a common subclass of bacterial polymers and are mostly present in the form of negatively charged sugar residues or as a result of the addition of negatively charged substituents. Uronic acids (e.g., GlcA) and nonulosonic acids (e.g., Neu5Ac, Sia) introduce negative charge into the RU, thereby rendering the polysaccharide a polyanionic polymer, much like the GAGs found in mammalian systems. Substituents on sugar residues such as pyruvate, phosphate and sulfate groups confer polyanionic character. The charged groups can be present both in the polysaccharide backbone and the side chains. Positively charged amines are sometimes present in the RU in conjunction with a negatively charged group. The CPSs from Neisseria meningitidis are representative for these types of polysaccharides. Types B and C are homopolymers of Neu5Ac and types W-135 and Y contain disaccharide RUs with Neu5Ac and a hexose.
Flexibility is a significant variable in bacterial polysaccharides (cf. the torsion angles φ and ψ). However, when the linkage is formed via O6, an additional degree of freedom becomes available at the glycosidic linkage (namely, the torsion angle ω) because of the exocyclic hydroxymethyl group in hexopyranoses. A (1-6)-linkage in the backbone of polysaccharides (Figure 3.4G) may result in a less rigid polymer with higher flexibility and random coil character. Likewise, the occurrence of this linkage in the side chains will make them more flexible. When furanose residues are part of the polymer, different ring conformations provide more options for introducing flexibility.
Cross-linking is another way that physical properties of bacterial polysaccharides can be varied. The cell wall of Gram-positive bacteria contains a particularly thick layer of peptidoglycan covalently cross-linked via short peptide sequences. Furthermore, teichoic acid polymers, built from glycerol or ribitol residues joined by phosphodiester linkages, are located within the cell wall. Mono- or disaccharide RUs containing amino sugars can be part of these repetitive structures, thereby giving rise to different cell wall teichoic acids. Polymers having a phosphodiester linkage in the backbone as part of the RU are referred to as “teichoic acid type.” CPSs of Haemophilus influenzae are built on this theme in which two (serotypes a and b) have RUs consisting of [ribitol-P-Hex-]n and two others (serotypes c and f) are made of [Hex-P-Hex-]n.
In summary, bacterial polysaccharides highlight the many ways in which diversity is built into oligo- and polysaccharides. Some diversity results from a larger set of sugar residues, some from branching, and some from modification with a wide variety of substituents, such as phosphate, sulfate, acyl, and amino groups. This diversity gives rise to different physical properties. It allows bacteria to mimic their hosts in an attempt to evade detection, and it provides a means of distinguishing self from competitive organisms. Structures discussed in this chapter are depicted in abbreviated IUPAC/IUBMB (International Union of Biochemistry and Molecular Biology) form in Table 3.1 for comparison to the common names, symbolic representations, and actual chemical structures used in the text and figures. This chapter serves as a preview of glycans discussed more thoroughly in the following chapters.
TABLE 3.1.
Oligosaccharides and polysaccharide repeating units
ACKNOWLEDGMENTS
The authors acknowledge contributions to previous versions of this chapter by Carolyn R. Bertozzi and David Rabuka and appreciate helpful comments and suggestions from Jenny Mortimer, Breeanna Urbanowicz, and Darón Freedberg.
FURTHER READING
- Laremore TN, Zhang F, Dordick JS, Liu J, Linhardt RJ. 2009. Recent progress and applications in glycosaminoglycan and heparin research. Curr Opin Chem Biol 13: 633–640. doi:10.1016/j.cbpa.2009.08.017 [PMC free article: PMC2787697] [PubMed: 19781979] [CrossRef]
- Zivkovic AM, German JB, Lebrilla CB, Mills DA. 2011. Human milk glycobiome and its impact on the infant gastrointestinal microbiota. Proc Natl Acad Sci 108: 4653–4658. doi:10.1073/pnas.1000083107 [PMC free article: PMC3063602] [PubMed: 20679197] [CrossRef]
- DeAngelis PL, Liu J, Linhardt RJ. 2013. Chemoenzymatic synthesis of glycosaminoglycans: re-creating, re-modeling, and re-designing nature's longest or most complex carbohydrate chains. Glycobiology 23: 764–777. doi:10.1093/glycob/cwt016 [PMC free article: PMC3671772] [PubMed: 23481097] [CrossRef]
- Cosgrove DJ. 2021. Re-constructing our models of cellulose and primary cell wall assembly. Curr Opin Plant Biol 22: 122–131. doi:10.1016/j.pbi.2014.11.001 [PMC free article: PMC4293254] [PubMed: 25460077] [CrossRef]
- Schmid J, Sieber V. 2015. Enzymatic transformations involved in the biosynthesis of microbial exo-polysaccharides based on the assembly of repeat units. Chembiochem 16: 1141–1147. doi:10.1002/cbic.201500035 [PubMed: 25873567] [CrossRef]
- Higel F, Seidl A, Soergel F, Friess W. 2016. N-glycosylation heterogeneity and the influence on structure, function and pharmacokinetics of monoclonal antibodies and Fc fusion proteins. Eur J Pharm Biopharm 100: 94–100. doi:10.1016/j.ejpb.2016.01.005 [PubMed: 26775146] [CrossRef]
- Moradali MF, Rehm BHA. 2020. Bacterial biopolymers: from pathogenesis to advanced materials. Nat Rev Microbiol 18: 195–210. doi:10.1038/s41579-019-0313-3 [PMC free article: PMC7223192] [PubMed: 31992873] [CrossRef]
- Whitfield C, Wear SS, Sande C. 2020. Assembly of bacterial capsular polysaccharides and exopolysaccharides. Annu Rev Microbiol 74: 521–543. doi:10.1146/annurev-micro-011420-075607 [PubMed: 32680453] [CrossRef]
- Review Oligosaccharides and Polysaccharides.[Essentials of Glycobiology. 2015]Review Oligosaccharides and Polysaccharides.Prestegard JH, Liu J, Widmalm G. Essentials of Glycobiology. 2015
- Review Recent advances in capillary electrophoresis separation of monosaccharides, oligosaccharides, and polysaccharides.[Electrophoresis. 2018]Review Recent advances in capillary electrophoresis separation of monosaccharides, oligosaccharides, and polysaccharides.Mantovani V, Galeotti F, Maccari F, Volpi N. Electrophoresis. 2018 Jan; 39(1):179-189. Epub 2017 Oct 9.
- Review Linkage Analysis of Oligosaccharides and Polysaccharides: A Tutorial.[Methods Mol Biol. 2021]Review Linkage Analysis of Oligosaccharides and Polysaccharides: A Tutorial.Black I, Heiss C, Carlson RW, Azadi P. Methods Mol Biol. 2021; 2271:249-271.
- POLYS 2.0: An open source software package for building three-dimensional structures of polysaccharides.[Biopolymers. 2014]POLYS 2.0: An open source software package for building three-dimensional structures of polysaccharides.Engelsen SB, Hansen PI, Pérez S. Biopolymers. 2014 Jul; 101(7):733-43.
- STUDIES ON HELICOSTYLUM PIRIFORME. II. UTILIZATION OF MONO-, OLIGO- AND POLYSACCHARIDES.[Mycopathol Mycol Appl. 1964]STUDIES ON HELICOSTYLUM PIRIFORME. II. UTILIZATION OF MONO-, OLIGO- AND POLYSACCHARIDES.MEHROTRA MD. Mycopathol Mycol Appl. 1964 Sep 30; 23:223-30.
- Oligosaccharides and Polysaccharides - Essentials of GlycobiologyOligosaccharides and Polysaccharides - Essentials of Glycobiology
Your browsing activity is empty.
Activity recording is turned off.
See more...
