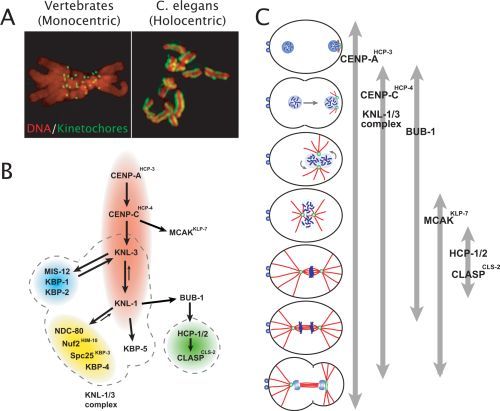
(A) Images comparing the localized kinetochores in a vertebrate tissue culture cell with monocentric chromosomes to the diffuse kinetochores in a C. elegans embryo with holocentric chromosomes. (B) Schematic illustrating the hierarchy for mitotic kinetochore assembly based on pair-wise depletion and localization assays, phenotypic analysis, and biochemical purifications. Dotted lines indicate groups of proteins that have been shown to co-purify from C. elegans extracts in immunoprecipitations and tagged protein isolations. The colored ovals group proteins together whose individual depletions result in a similar phenotype. (Red) "Kinetochore Null"/KNL proteins whose depletion is characterized by a failure to assemble kinetochores that are competent to form spindle microtubule attachments. Consequently, segregation is severely defective and spindle poles separate prematurely. (Blue) "MIS" proteins whose depletion results in relatively subtle chromosome segregation defects. In MIS embryos, kinetochore assembly occurs but at a slower rate and to a reduced extent relative to wild-type. (Yellow) "NDC" proteins whose depletion results in a chromosome alignment and segregation defects of intermediate severity, relative to the KNL and MIS classes. In NDC embryos, attachments that can sustain tension fail to form. Consequently, spindle poles separate prematurely. (Green) HCP/CLASP proteins, whose depletion causes sister chromatids to co-segregate to the same spindle pole. This defect likely arises from an inability to polymerize microtubules at kinetochores. (C) Schematic illustrating the temporal window during the first mitotic division when each of the indicated proteins localizes to kinetochores. Figure and images courtesy of Susan Kline-Smith and Arshad Desai, respectively.