- Exon count:
- 8
| Annotation release |
Status |
Assembly |
Chr
|
Location |
| RS_2024_08 |
current |
GRCh38.p14 (GCF_000001405.40) |
20 |
NC_000020.11 (63895126..63936011) |
| RS_2024_08 |
current |
T2T-CHM13v2.0 (GCF_009914755.1) |
20 |
NC_060944.1 (65714973..65755860) |
| RS_2024_09 |
previous assembly |
GRCh37.p13 (GCF_000001405.25) |
20 |
NC_000020.10 (62526479..62567364) |
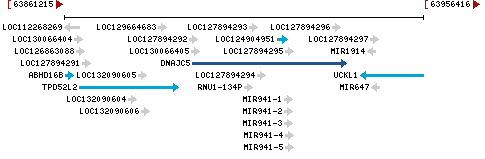
Chromosome 20 - NC_000020.11